RAS instructions to editors
You have access to edit all the information in the system only within the taxonomic group assigned to you as an associate editor. Before you change existing information, it is important that you do not "overwrite" valuable information. For example, a common misspelling of a name can be valuable to be kept in the database. In that case, it is better that you create a new entry and link the misspelling to the correct name. Below you will find a list and a short explanation of what is possible in this system. If you have any questions, please do not hesitate to ask the Database Management Team (DMT) by clicking here
Published as Horton, et al. (2017). Improving nomenclatural consistency: a decade of experience in the World Register of Marine Species. European Journal of Taxonomy. 389: 1-24. doi: 10.5852/ejt.2017.389What information do we prioritize?
| Required | Classification, species name, authority, publication year and one habitat flag |
| Highly desired | Basionym, original publication reference, holotype information (type locality, museum, number, ...) |
| Optional | additional synonyms, references, images, morphological description, specimens, distribution, attributes, weblinks |
Instructions
- Edit taxon details
- Authority format
- Add a new accepted taxon
- Add a new unaccepted taxon
- Add a new environment flag
- Add a fossil flag
- Add a new source
- Link a source to a taxon
- Add a new vernacular
- Add a geographical location
- Add a distribution record from a picklist
- Add a distribution record on a map (distribution map entry tool)
- Add many distribution records more easily
- Add a new specimen
- Add a feeding type to a taxon
- Add a link to a taxon
- Add a PDF to a taxon
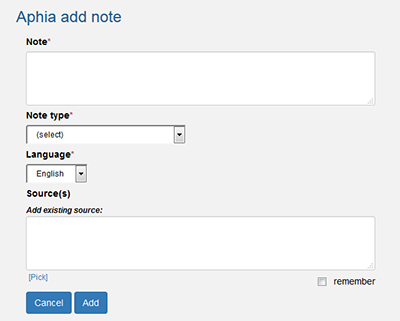
- Add a note (characteristic) to a taxon
- Add an image to a taxon
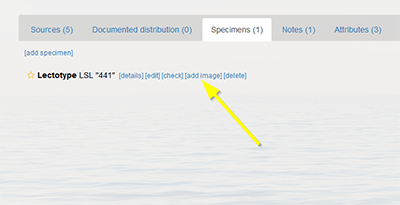
- Add an image to a specimen
- Add a new IDkey
- Add/remove a taxon to an IDkey
- Change a taxon name
- Change the status of a name
- Change the record status
- Change (the status of) a distribution
- Change collection information for a specimen
- Change a source
Frequently asked questions
- How come there is a difference between RAS and some of the contexts like Isopoda etc ?
- How can I know what species of my taxon are in RAS, but are not part of my collection?
- How do I know what species have been added or changed by my co-editor?
- How can I update the classification?
- How can I add a subspecies when the species name on itself is not in use?
- Can I link an unaccepted name to more than one taxon?
- How do I enter an occurrence record in a paper that mentions the original paper, which also used a synonym name?
- What should I do if not all genera can be assigned to a subfamily for example?
- Can we emphasize preferred vernacular names?
- Can I add taxa that are in press?
- Can you explain how the citation on each page works?
- When adding a specimen, should I use the original species name or the currently accepted?
- How are we dealing with the correct citation of the year of publication of a new species´ description when the online version and the printed version appear in different years?
- How should I handle taxa that were originally introduced with a question mark: e.g. "?Bifrontia zanclaea"?
- How should I handle species that currently are *tentatively* placed in synonymy (the kind of thing that is usually preceded by a "?" in synonymies)?
- How can I export a source in RIS and BibTex format?
- What does the † stand for in e.g. Castexia douvillei Mercier, 1936 † ?
- How can I download the results of my taxon search?
- What literature pdfs are openly available?
- Where should I place unaccepted genera in the taxonomic hierarchy?
- How to deal with unavailable names?
- How to deal with species with accepted subgenera using the status of alternative representation?
Please ask the data manager for a password if you should not yet have received one. The access rights are linked to a higher taxa in the classification tree. In other words, you will only be able to edit information in your particular taxon group of expertise.
If you have a separate web interface (e.g. https://www.marinespecies.org/copepoda), then always log in via this website,
so that all the data you entered/edited is automatically part of this collection.
You can always go to [My Aphia] to check your edit rights and change your password or other settings if needed. You can reach 'My Aphia' by clicking your name in the upper right corner.
Top Edit taxon details If you want to change the details of a taxon that already exists in RAS, you search the taxon and click on [edit taxon] to the right of the taxon. You can change all the information except the AphiaID.
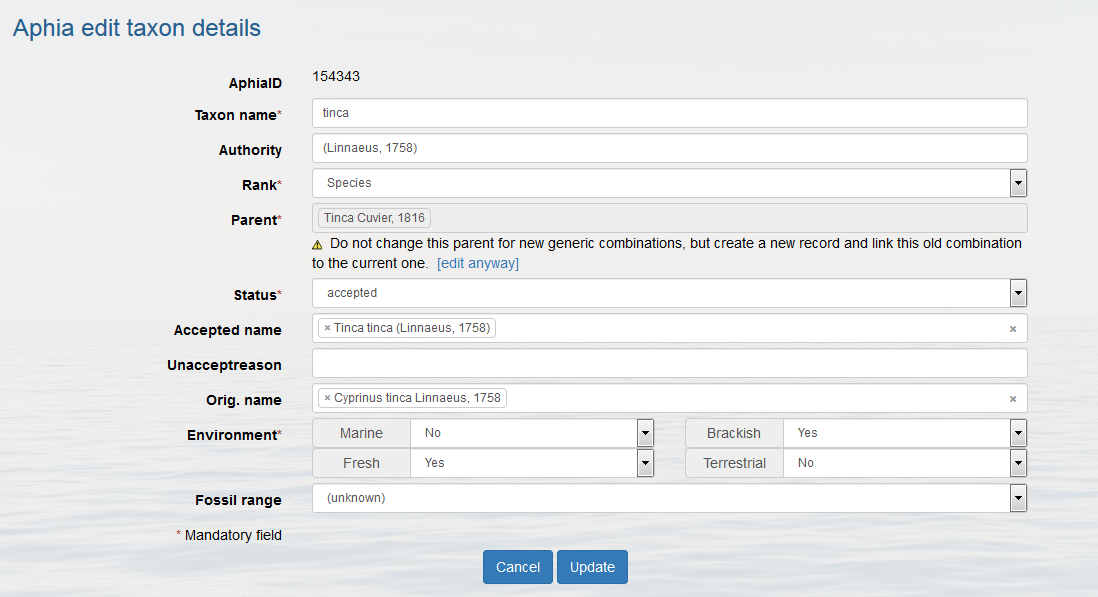
The authority format differs, depending to which kingdom the species belongs.
The rules are laid out in different Codes: the International Code of Zoological Nomenclature (ICZN), the International Code of Nomenclature for algae, fungi, and plants (IAPT) (previously called the International Code of Botanical Normenclature - ICBN), the International Code of Nomenclature of Bacteria (ICNB), the International Code of Nomenclature of Prokaryotes and the International Code on Taxonomy of Viruses (ICTV).
- In general, the following format is used in the database:
- For original names:
- the first part is(are) the name(s) of the author, separated by a comma and space.
Example: Linnaeus, 1758 - In case of multiple authors, an "&" is place right before the last author. This is then followed by a comma, space and the year of the publication.
Example: Dózsa-Farkas, Felföldi, Nagy & Hong, 2018
- the first part is(are) the name(s) of the author, separated by a comma and space.
- For new combination names:
the authority of the original name must be placed between brackets.
Example: (Lacepède, 1802) - Year of publication
It is recommended to always include the year of publication, as this helps in e.g. the compilations of the number of species described per year
- For original names:
Examples typically come from parent and sibling(s) all active in taxonomy or husband and wife both active taxonomists.
. E.g. C. Monniot & F. Monniot (Ascidiacea taxonomists); H. Adams & A. Adams (Mollusca taxonomists)
The correct use of brackets is extremely important: this indicates if the name is an original name or not.
E.g. Dicentrarchus labrax (Linnaeus, 1758), which is an new combination. This is clear because of the brackets around the authority of the original name, which is Perca labrax Linnaeus, 1758. So historically, Perca labrax was the first name published for the European seabass, described and published by Linnaeus in 1758. Later on, this species was moved to the genus, Dicentrarchus, which results in a new combination, but keeping the original authority between brackets.
For taxa described under the Zoological Code: Where the authority of a taxon differs from the author(s) listed on a publication, editors should use the following format:
‘TaxonAuthors in PublicationAuthors, Date’. If PublicationAuthors contains 3 or more authors, the authors may be expressed by use of the term "et al." following the name of the
first author, provided that the original description is linked. For example, "Cuniculomaera Tandberg & Jażdżewska in Brandt, Chen, Engel, Esquete, Horton, Jażdżewska, Johannsen,
Kaiser, Kihara, Knauber, Kniesz, Landschoff, Lörz, Machado, Martínez-Muñoz, Riehl, Serpell-Stevens, Sigwart, Tandberg, Tato, Tsuda, Vončina, Watanabe, Wenz & Williams, 2024"
may be shortened to "Cuniculomaera Tandberg & Jażdżewska in Brandt et al., 2024".
Top
Add a new accepted taxon
If you do a search for a taxon and do not find it, then you can add it. Every name
in RAS
is linked to a "parent" taxon: e.g. a species epithet is linked to its genus or subgenus, a genus is linked
to its family or subfamily. Search for the parent taxon using the search tool and click on [add child taxon].
This will open a new window "Aphia add a taxon", with an empty taxon form. Fill in all the necessary information
and click on [add]. The fields indicated with a red asterisk are mandatory fields. The information in the parent
field cannot be changed, as you indicated that you wanted to create a "child" taxon for the taxon you started
from. The record status of the name will automatically be "checked by editor" and in the edit history at the
bottom of the page, the "date", and created by "your name" will be listed. When adding the authority make sure
you use author(s), year and brackets () and/or accents if needed. Select at least one environment
(marine/brackish/freshwater/terrestrial) if you create a new taxon.
Possible accepted statuses for a name:
- Accepted: Valid name (ICZN) or name considered to be taxonomically correct (ICBN)
- Unreplaced junior homonym: To indicate the unreplaced invalid status of the junior, or later established name, or in the case of simultaneous establishment, the name not given precedence by a first reviser or by an ICZN or ICN ruling.
See also: How can I add a subspecies when the species name on itself is not in use?
See also: Can I add taxa that are in press?
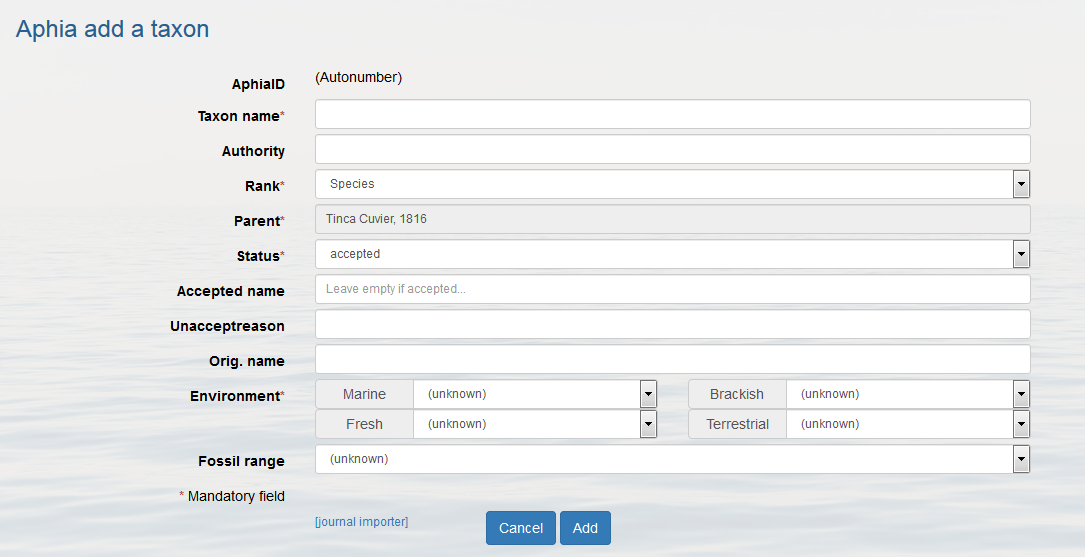
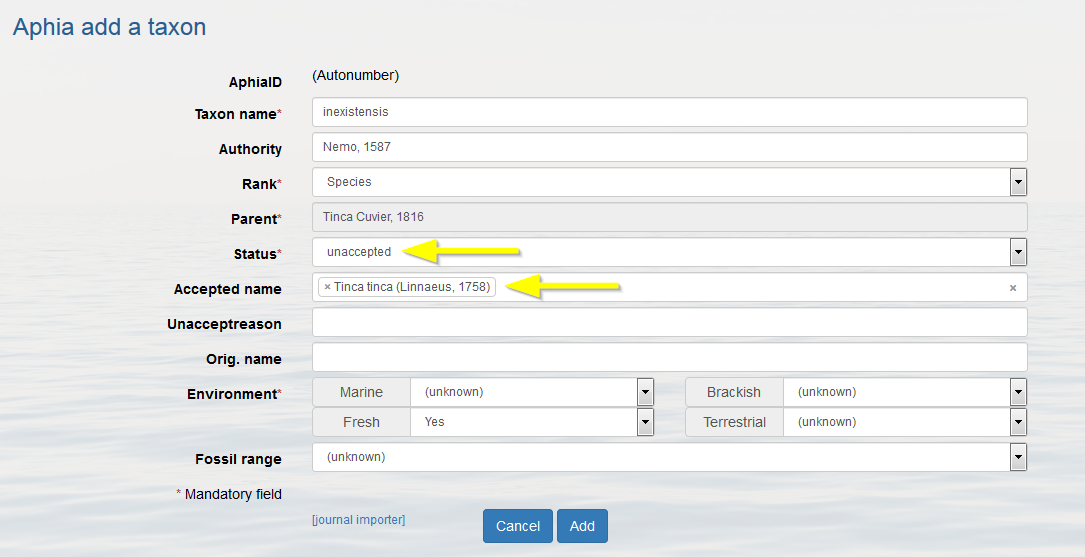
Go to the parent taxon and follow the same procedure as for adding a new accepted taxon (click on [add child taxon]). Now change the status into "unaccepted" and enter the "accepted name", this is a drop-down list. If the accepted name is not found, then add it first. If possible fill in all other information and click on [add].
In RAS invalid (unaccepted) synonyms are linked to accepted taxa. This requirement is not needed for cases like:
- nomen nudum: i.e. a name that does not comply with the name requirements of the codes, such as lack of a description or diagnoses or reference to a description or diagnosis or a type specimen is lacking for publications after 1999
- nomen dubium: i.e. a name of uncertain application, because it is not possible to establish the taxon to which it should be referred. A good example is the "Ascothoracida" genus Laocoon. There is a debate whether this is based on a parasite or on a detached piece of the host. It is clearly a dubious name
- temporary name: e.g. to create higher rank taxa to accommodate child taxa for which the classification is not sorted yet
- species inquirenda: i.e. an incompletely defined species that requires further characterization, it is impossible to identify the species
- incertae sedis: of an uncertain seat; i.e. of uncertain taxonomic position; or when the original material has not been studied or the description is insufficient. Usually applied to family and generic names
Possible unaccepted statuses for a name:
- Unaccepted: Synonym name, or anything that is not accepted
- Nomen nudum: A Latin term referring to a name that, if published before 1931, fails to conform to Article 12; or, if published after 1930, fails to conform to Article 13. A nomen nudum is not an available name, and therefore the same name may be made available later for the same or a different concept; in such a case it would take authorship and date [Arts. 50, 21] from that act of establishment, not from any earlier publication as a nomen nudum.
- Interim unpublished: An as yet unavailable name (until in a print issue) which has been published online only, in a work that does not show evidence of ZooBank registration (ICZN Article 8.5)
- Superseded combination: When a species is transferred to another genus, and is also known as a new combination
- Junior homonym: To indicate the invalid status of the junior, or later established name, or in the case of simultaneous establishment, the name not given precedence by a first reviser or by an ICZN or ICN ruling. This is known as a later homonym in the ICN.
- Junior subjective synonym: Applied as the reason for the unaccepted status for the junior, or later established, subjective synonym (a published opinion that two names apply for the same taxon). This is known as an heterotypic synonym in the ICN.
- Junior objective synonym: Applied to the later established name in cases where two or more scientific names have the same type material. This is known as a homotypic synonym in the ICN.
- Nomen oblitum: A Latin term (meaning ‘forgotten name’) applied after 1 January 2000 to a name, unused since 1899, which as a result of an action taken under ICZN Article 23.9.2 does not take precedence over a younger synonym or homonym in prevailing usage
- Misspelling - incorrect original spelling: An error in the name's original publication that doesn't comply with ICZN naming rules. Corrections to these errors are termed "justified emendations" by the ICZN, requiring compliance with ICZN rules and guidelines (see section 8 in Welter-Schultes 2012)
- Misspelling - incorrect subsequent spelling: Erroneous name in the literature that need to be captured, e.g., when widespread, or needed in the database
- Unjustified emendation: Nomenclatural act under ICZN article 33.2 and must strictly be a ‘demonstrably intentional change’ of the original spelling
- Incorrect grammatical agreement of specific epithet: Mandatory change in spelling consequent upon changes in rank or combination. See Welter-Schultes (2012) section 8.1.6
- Misapplication: When a name has been misapplied to an incorrectly identified specimen.
-
Unavailable name: Not compliant with the relevant ICZN or ICN code
See also How to deal with unavailable names? - Superseded rank: When a subspecies, subgenus, subfamily, etc. is moved up or down within the hierarchy (i.e., subspecies now accepted as a species or vice versa)
- Nomen rejiciendum: (1) A name which, under the provisions of the Code, cannot be used as a valid name and which is set aside in favour of another name. (2) A name which, as a matter of taxonomic judgment, is either treated as a junior subjective synonym (q.v.) of a name used as valid or is believed not to be applicable to the taxon under consideration.
More information about name statuses can be found at Hawksworth, D.L. 2010. Terms Used in Bionomenclature. The naming of organisms (and plant communities). Copenhagen: Global Biodiversity Information Facility, 216 pp
See also: Can I link an unaccepted name to more than one taxon?
You can specify the fossil flag while creating a new taxon or you can add a fossil flag to an already existing taxon. In the latter case, go to the taxon page and click on [Edit taxon].
There are 4 fossil flags:
- (unknown): this is the default value and is used when the fossil state is unknown
- recent only: used to define taxa that are known from only recent, non-fossil, records
- fossil only: used to define taxa that are known from only fossil records
- recent + fossil: used to define taxa that are known from both non-fossil and fossil records
Possible search options, will return:
- (any): all taxa, independent of their fossil range
- (unknown): all taxa of which the fossil range is only unknown
- recent only: all taxa that are known from only recent, non-fossil, records (this will exclude taxa with the unknown flag)
- fossil only: all taxa that are known from only fossil records
- recent + fossil: all taxa that are known from both non-fossil and fossil records
- extant, not fossil-only: all taxa that are known from: (1) both non-fossil and fossil records, (2) only recent records, and (3) with an unknown fossil range. In other words: taxa only known from fossil records are excluded.
Log in to the database, go to "Literature" in the menu, and click on [Add source] on the bottom of the page.
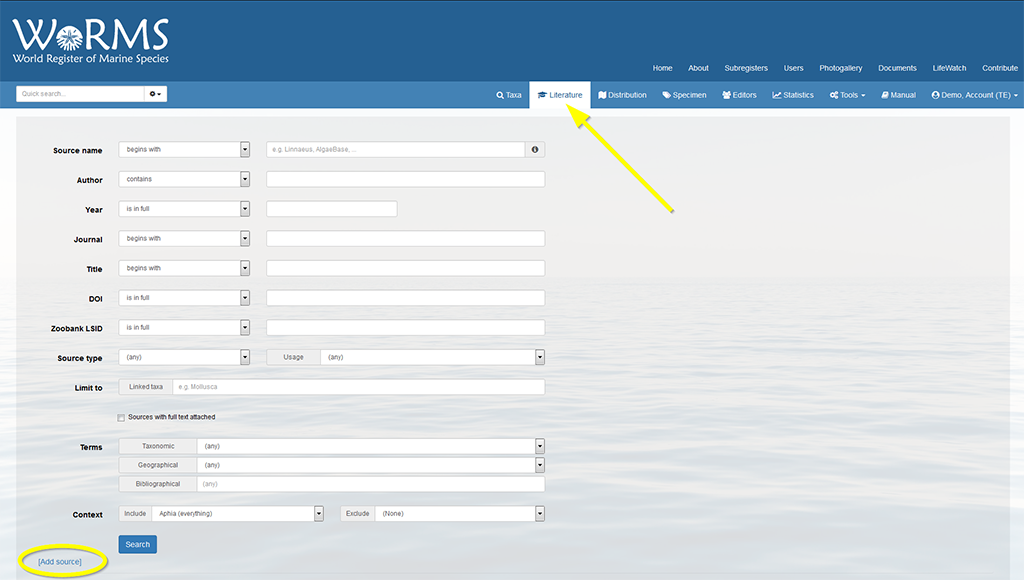
Instead of storing the source in just one field, sources can now be stored more efficiently, in a so-called "atomized" way. The "Source name" field is split up into 6 fields: Digital Object Identifier (DOI), author, year, title, journal and suffix (volume, pages, etc.). When entering new sources, these fields can be filled in manually, but sources can also be completed and/or atomized automatically when the DOI, ZooBank LSID or full citation is available.
Paste the DOI, ZooBank LSID or full citation in the first field. Make sure the "auto" checkbox is checked. If not, you can also click on "Parse".
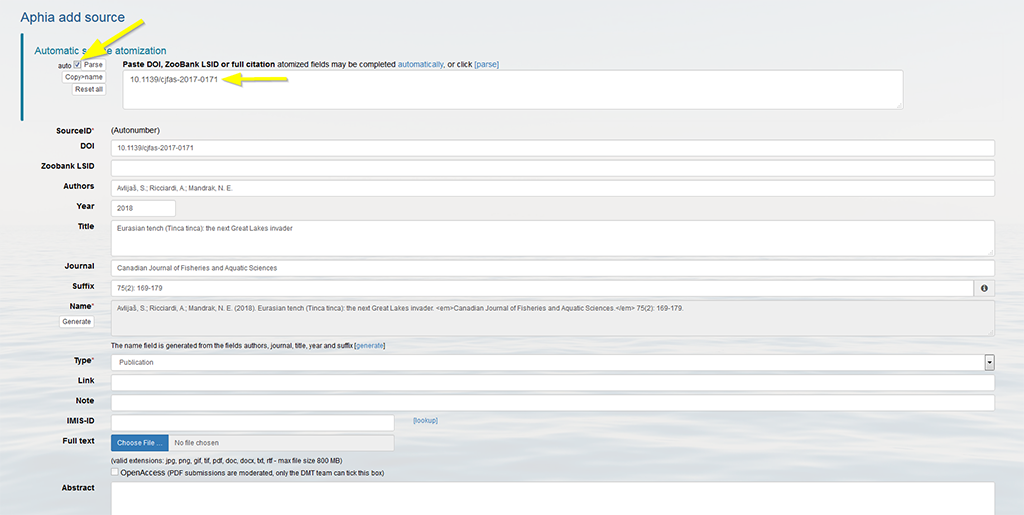
The other fields are filled in automatically. Make sure the parsing of the fields was done correctly; a manual check is highly recommended to verify the automatically parsed data. If mistakes would have occurred, you can edit the fields manually. In case of manual changes, don’t forget to click on "Generate", to complete the source name field.
It is also possible to add additional information (note, IMIS-ID, abstract, etc.), and to upload the full text publication. The maximum file size is set at 800MB. If you would like to add many sources (bulk upload), upload a larger file or if you would experience any problems during the uploading, please contact us.
If all fields are filled in correctly, click "Add", and the atomized source will be added to the database.
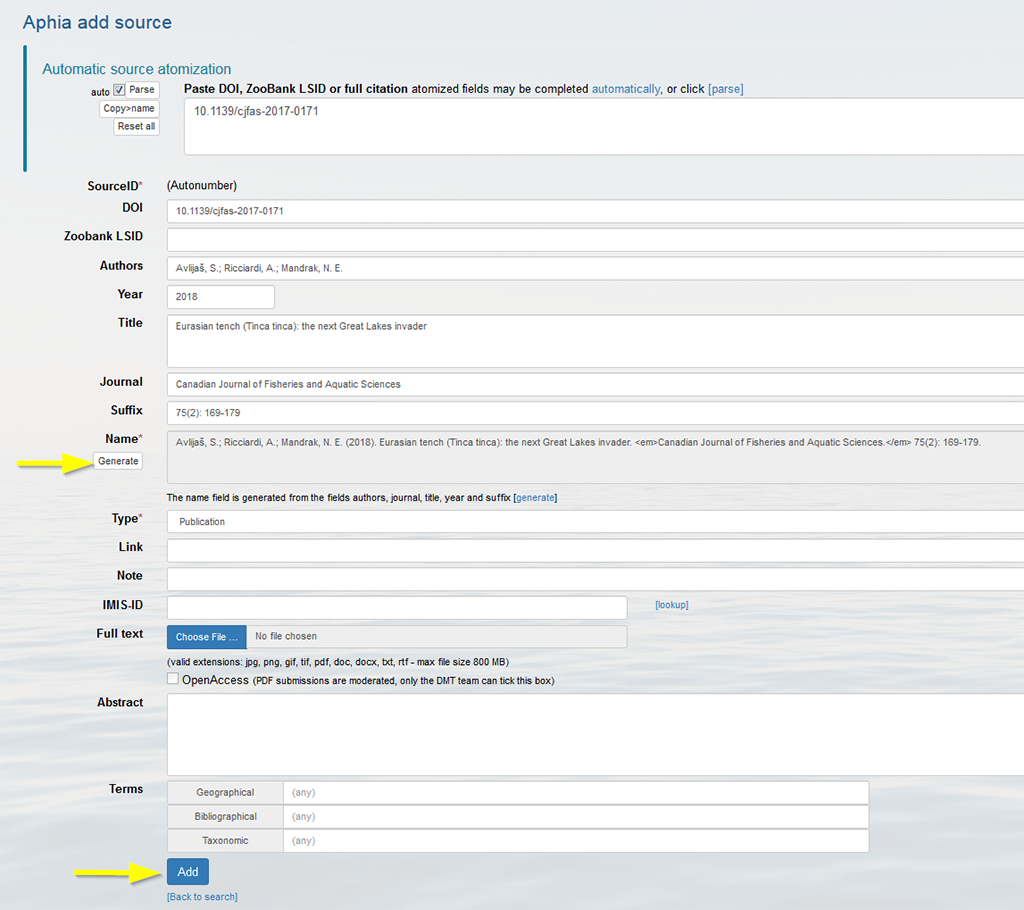
The automatic atomization of sources is based on external services: CrossRef, ReFindit and Grobid. More information can be found here.
We highly recommend to use the automatic parsing and atomization of sources (i.e. if the DOI or ZooBank LSID is available), since this significantly reduces the risk of having duplicate sources in the database.
See also: How can I export a source in RIS and BibTex format?
Top Link a source to a taxonTo link a new source, go to the taxon details page. Before you link a source to a taxon you have to see if it isn’t already linked.
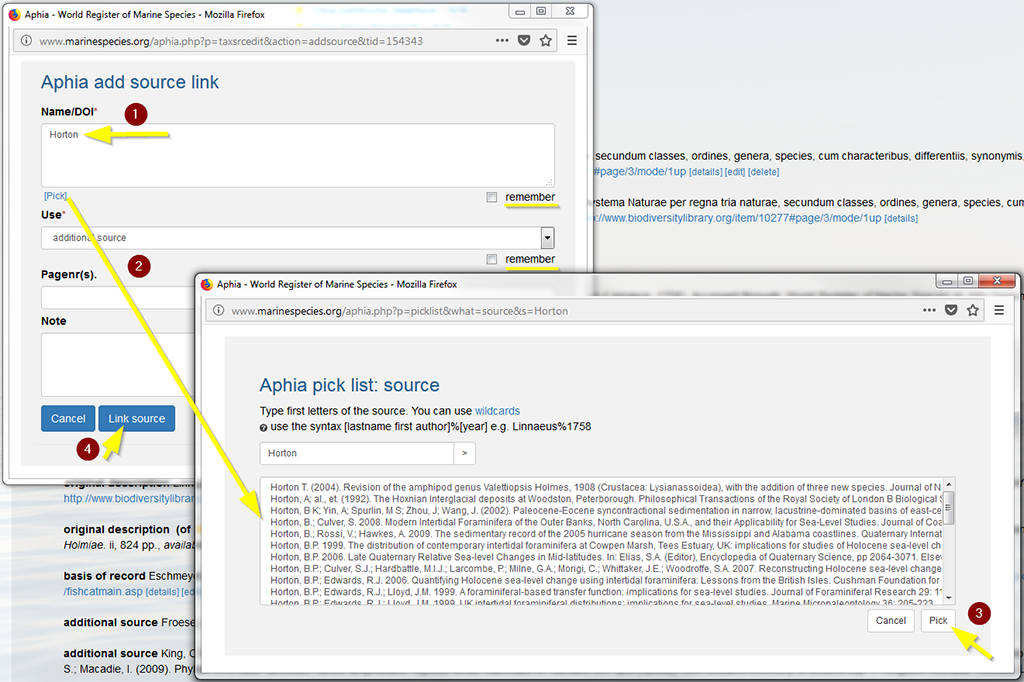
Click on [link source]. This will open a new window "Aphia link a source to a taxon", with an empty form. Enter the first letters of the first authors’ last name and click on [pick]. This will open a new window "Aphia pick list: source", where you can select your source from the drop-down list and click on [pick]. Fill in the Use and Pagnr. and click on [link source]. If this is for a new species, then pick the correct source, change 'Use' to original description and add the pages that the species was described on.
You can also search for an author that is not first author by using the wildcard % and last name to find anywhere in the author string. You can select the "remember" button, so that the reference automatically appears the next time you want to link a source to a taxon (helps a lot if you enter many names of a single publication). The system is set so that the accent marks used in author names will not hamper the search (e.g. if you type in Gomez you will get Gómez or Mollmann will get Möllmann).
You can add multiple sources if needed. Say you want to record all the authors that published on the taxa, then you create a new link for each source as needed. And then change the 'Use' to one of the 14 choices (see below) as necessary.
Source "use" values:
- Original description: publication in which the name is made available.
- by a description in words (mandatory after 1930, except for replacement names), i.e. its basionym, or
- by a figure and a name without a description (tolerated before 1931), or
- by reference to a description published elsewhere (also tolerated before 1931, ICZN art. 12.2.1), or
- by replacement (ICZN art. 12.2.3)
- Basis of record: source consulted when entering the present record
- Additional source: source that contains additional information about the present taxon
- Source of synonymy: source that contains information on the synonymized taxa
- Redescription: source in which the present taxon is redescribed in a modern way
- New combination reference: source in which the present combination was first proposed
- Status source: source that contains information on the current status of the present taxon
- Toxicology source: source that contains information on the toxic nature of the present taxon
- Taxonomy source: source that contains information about the classification of the present taxon
- Ecology source: source that gives information about the relations and interactions between the present taxon and its environment, including other organisms
- Identification resource: source that contains information on the identification of the present taxon
- Subsequent type designation: publication in which the type-species or –specimen of the present taxon was designated
- Misapplication: source in which the present taxon name was used in an erroneous way
- Context source: source used to link a taxon to a context (subcollection) of Aphia
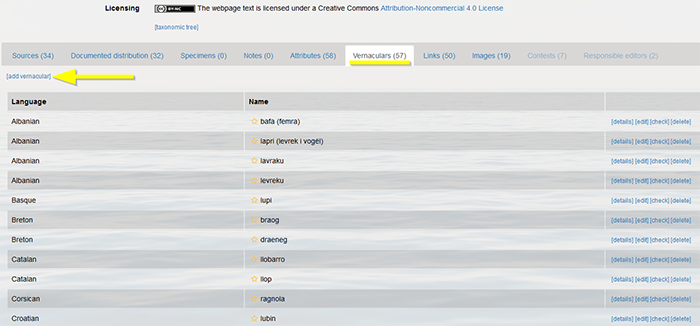
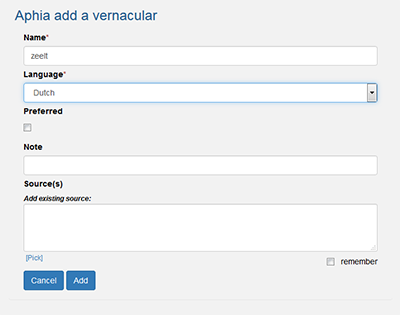
Vernaculars can only be written with a capital if they refer to nouns, geographical, German or Japanese names. Otherwise, they should always be written without a capital.
See also: Can we emphasize preferred vernacular names?
Go to the Marine Regions Gazetteer, login with your RAS password and go to [My Marine Gazetteer].
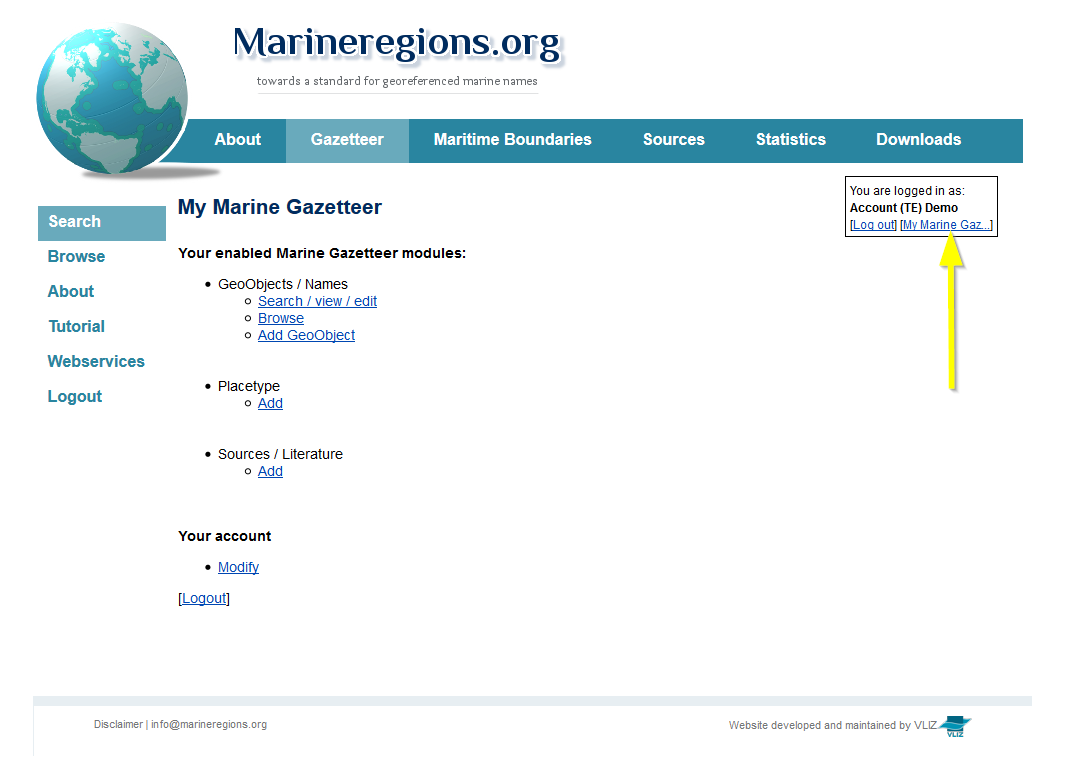
It is possible that the location is not listed in the drop-down list in RAS, but is present in the gazetteer. First search if the location is available in the gazetteer (via the search tool), before you start adding a new location record. If the location is present in the gazetteer, then click on [add to aphia] and the location is added to the drop-down list in RAS.
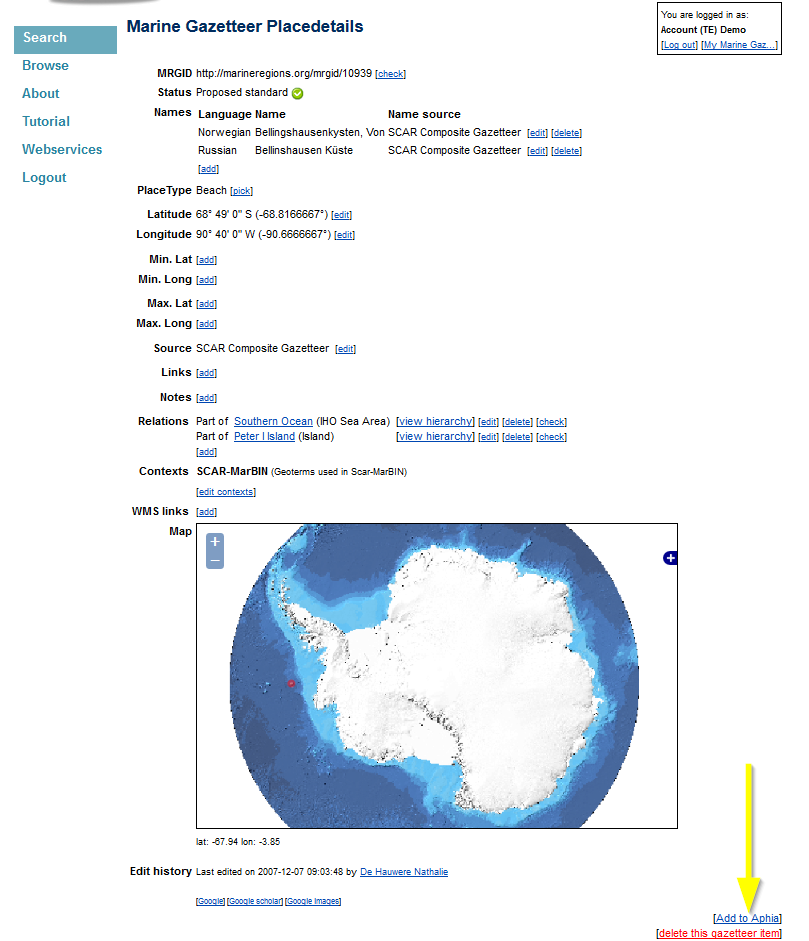
If the location is also not present in the gazetteer, than click on the add buttons to add a geographical location, source or place type (find the buttons on [My Marine Gazetteer]). Fill in the necessary fields and click on [create]. A link to the parent location will create the hierarchical tree of the localities. For marine locations (such as lagoons, bays, sandbanks, etc… use the country name + Exclusive Economic Zone (or the sea/marine area it belongs to) as its parent, instead of the country name, which refers to the land part. The geographical location is automatically transferred to RAS.

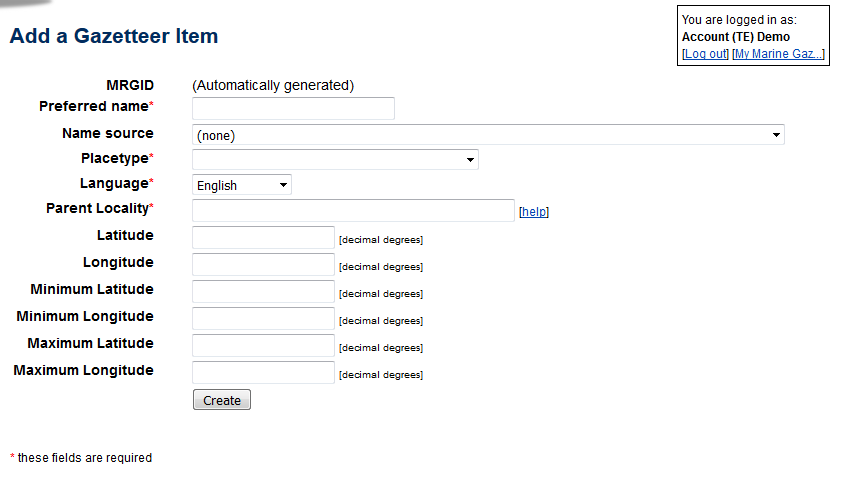
Go to the bottom of the taxon page, click the tab 'Documented distribution' and click on [add distribution] in the upper left corner. This will open a new window "Aphia add distribution record".
Pick the geographical location as detailed as possible from the picklist. If the place name is not in the list, add it first (see above). Do not enter e.g. "Oostende, Belgium", but only enter "Oostende". Use the country name Exclusive Economic Zone if it is a marine species, rather than just the country name. The geographical names are linked to the Marine Regions gazetteer, which hierarchically structures the geo-names.
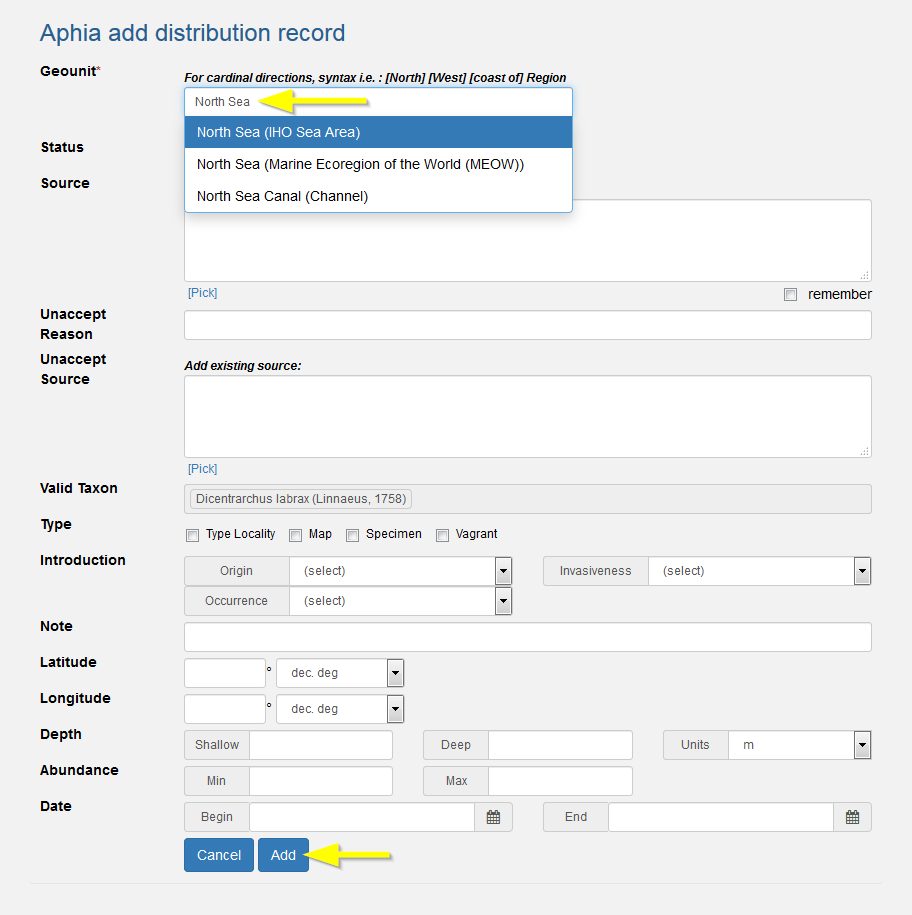
There are 3 possibilities for the status of a distribution record:
- Valid: the record is certainly valid
- Doubtful: doubts on whether the species really lives there
- Inaccurate: taxon was cited for this area but it is clear that it does not live there (misidentification, wrong label, etc.)
In the field "Source" you can add the source for the taxon distribution.
You can also add the reason ("Unaccept Reason") and source ("Unaccept Source") for doubtful or inaccurate distributions.
In case of introduced species, you can indicate the Origin; Invasiveness and Occurrence of the taxon. For more information about the terminology used to describe introduced taxa, go to the World Register of Introduced Marine Species (WRIMS).
Latitude and Longitude are only needed for point locations. In this case the geo-unit will be a general place name (e.g. seas, countries, etc.). You can choose between the decimal or DMS (Degrees, Minutes, Seconds) format, click on DMS to switch. For decimals, do not forget to use the minus sign for South and West. For example 40S = -40.
The date should be the begin date. The end date can only be filled in if the observation was only made during a certain period.
Note that you can enter the type locality either via the specimen module or as a distribution (flagging on "type locality"). If the type locality is entered in the specimen module, then there is no need to enter it again as a distribution record (because this is something we can create automatically). Some explanation of the various flags linked to distribution records:
- Type locality: the locality of the holotype
- Map: derived from indication on a map of the place where the species was found
- Specimen: this record represents a specimen deposited in a museum
- Vagrant: species appeared accidentally outside its natural range
If all the necessary geo units are selected and all fields are filled in, click on "Add".
See also: Change (the status of) a distribution
Top Add a distribution record on a map (distribution map entry tool) Go to the bottom of the taxon page, click the tab 'Documented distribution' and click on [add map distribution(s)] on the upper left corner. This will open a new window "Aphia add distribution record". Select an appropriate (standard) layer. For more information about the different geounits, go to the Marine Regions gazetteer.Select one or more geo units on the map.
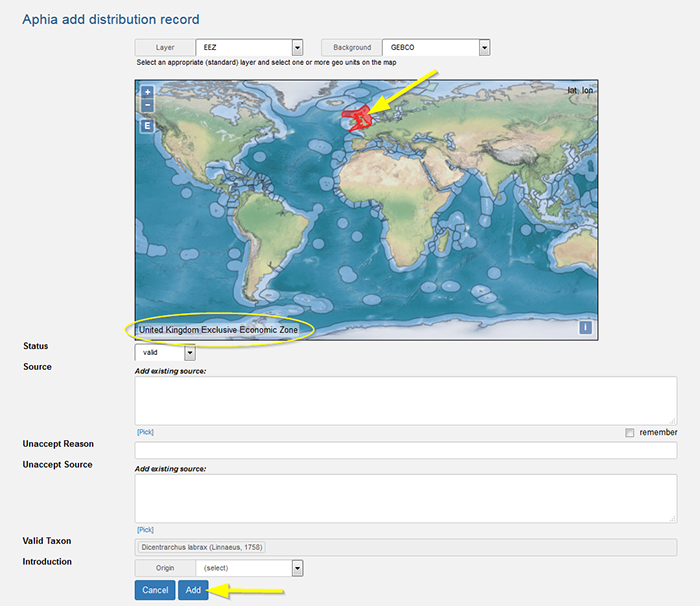
There are 3 possibilities for the status of a distribution record:
- Valid: the record is certainly valid
- Doubtful: doubts on whether the species really lives there
- Inaccurate: taxon was cited for this area but it is clear that is does not live there (misidentification, wrong label, etc.).
In the field "Source" you can add the source for the taxon distribution.
You can also add the reason ("Unaccept Reason") and source ("Unaccept Source") for doubtful or inaccurate distributions.
In case of introduced species, you can indicate the origin of the taxon: Alien, Native, Native – Endemic, Native - Non-endemic, Origin uncertain, Origin unknown. For more information about the terminology used to describe the origin of a taxon, go to the World Register of Introduced Marine Species (WRIMS).
If all the necessary geo units are selected and all information is filled in, click on "Add".
See also: Change (the status of) a distribution
Top Add many distribution records more easily When you have a species list of a single location or a list of geographical locations of a single species, then it is much easier to enter these distribution records using the "rapid distribution entry" page. Go to the [My Aphia] screen and click on "Rapid Distribution Entry". You can ask the system to remember the taxon or the geographical location for the next entries. This saves you time because you don't need to select these items over and over again. Also the source and other information that you add to the fields is automatically remembered by the system.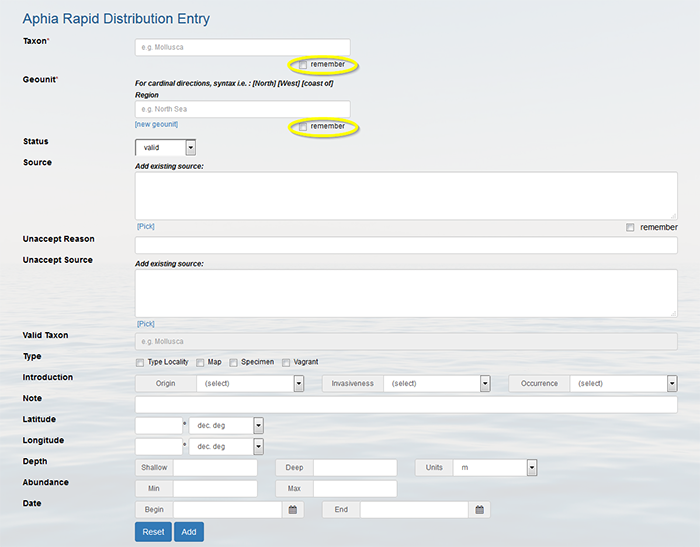
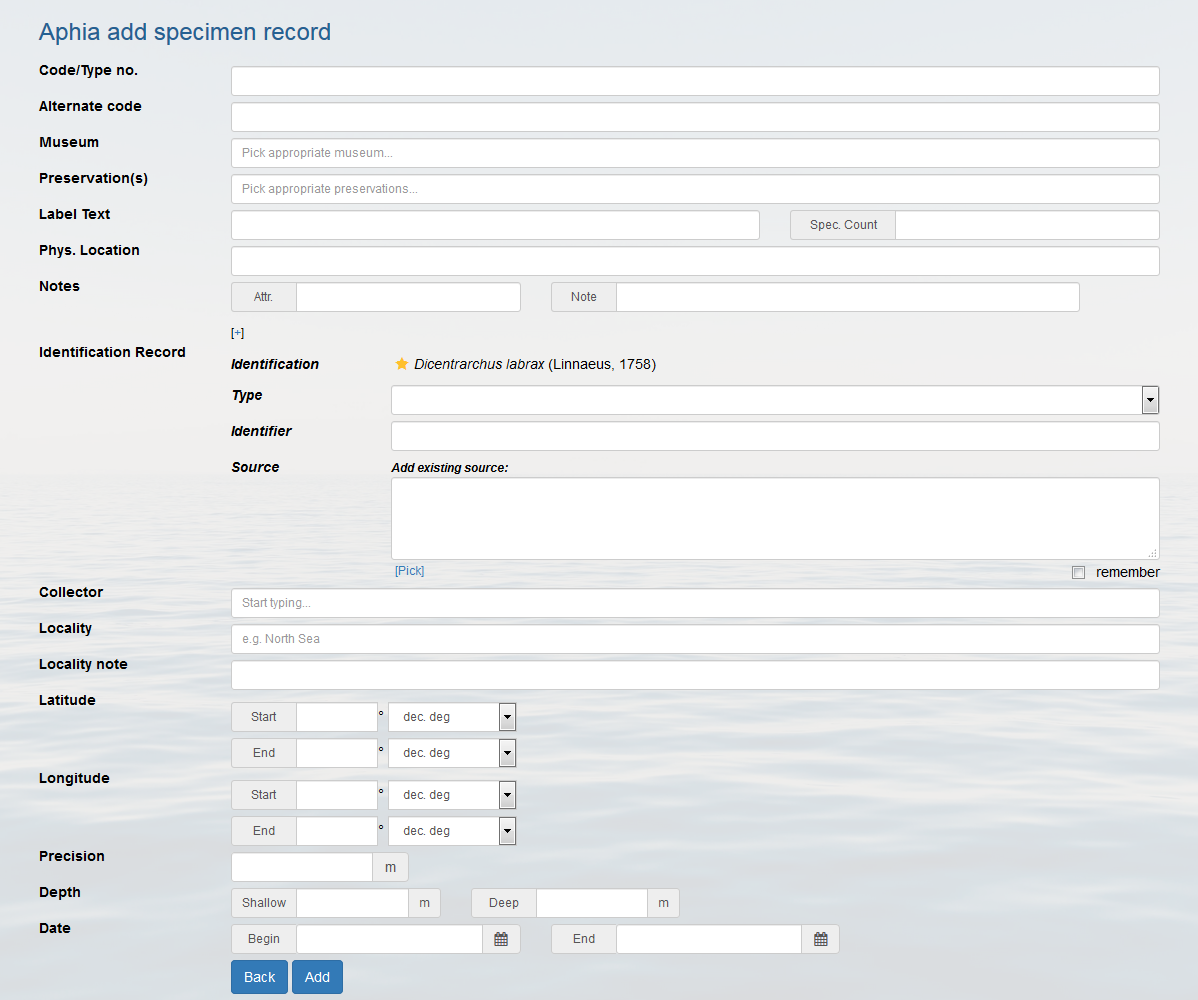
See also: When adding a specimen, should I use the original species name or the currently accepted?
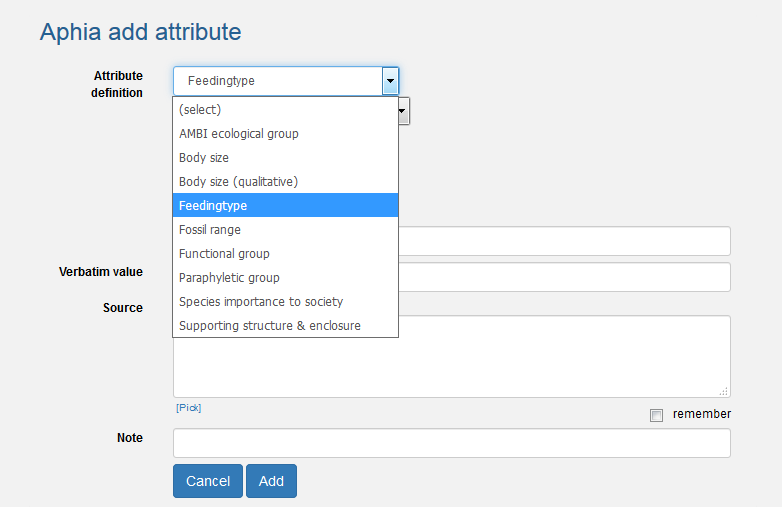
This is where you can add info about the taxon, how and where it feeds. Go to the bottom of the taxon page, click the tab 'Attributes' and click on [add attribute] in the upper left corner. This will open a new window "Aphia add attribute". Next to Attribute definition, select 'Feedingtype' from the predefined drop-down list. Select the type of feeding, add more information if possible and click on [add]. Notice there is also a drop-down list ‘Stage’: if this feeding type only occurs in a certain life stage of the taxon you can indicate this here. You can add a feeding type for every life stage if necessary. You can also add host information for parasites and symbionts. There is field to fill in the host. There is also a note field to add information such as where on the host it lives (e.g. collected from nasal cavity or attached to gills, etc…). If RAS does not find a match in the host name, then it means it is not in the database and has to be added. Contact us here, and we will add the host taxon name.
Possible feeding types:
- Carnivore: flesh-eating organism
- Commensal: an organism participating in a symbiotic relationship in which one species derives some benefit while the other is unaffected
- Deposit feeder: feed on organic material contained in the soft sediments of the sea bottom. Sometimes called ‘bottom-feeder’
- Deposit feeder: non-selective: eats any kind of deposits
- Deposit feeder: selective: prefers a certain type or size of particles
- Deposit feeder: subsurface: exploit food located deeper in the sediment (few cm)
- Deposit feeder: surface: feed on particles on the surface
- Detritus feeder: animals that consume decomposing particulate organic matter (as opposed to dissolved organic matter)
- Epigrowth feeder: eating from the top layer of plant and algae growth
- Filter feeder: organism that feeds on particles of organic matter that are suspended in water by creating a water current
- Grazer: feeds on plants and algae by scraping along a surface
- Herbivore: plant eating organism
- Interface grazer: eating from the middle layer of plant and algae growth
- Not feeding: doesn’t eat
- Omnivore: all-eater or polyphage, eats plants, animals, algae, fungi, …
- Parasite: an organism that grows, feeds, and is sheltered on or in a different organism while contributing nothing to the survival of its host
- Parasitic: ectoparasitic: shelters on the host
- Parasitic: endocommensal: commensalism is an interspecific, symbiotic relationship in which two different species are associated, wherein one is benefited and the other neither benefited nor harmed. Here the beneficial organism lives in the other organism
- Parasitic: endoparasitic: shelters in the host
- Predator: an organism that preys on other animals as a source of food
- Predator/omnivore: an organism that actively searches for food but eats everything
- Scavenger: organism that feeds on dead animal material, and sometimes also drift algae
- Suspension feeder: organism that feeds on particles of organic matter that are suspended in water
- Suspension feeder: facultative: organism that can choose between suspension feeding or another type of feeding
- Symbiotic: epizoic: Epizoic mean that one organism is living on the surface of another organism. The relation is beneficial for at least one member
- Symbiotic: unspecified type: a close, prolonged association between two or more different organisms of different species that may, but does not necessarily, benefit each member
- Unknown
Possible life stages:
- Egg: animal develops in the female gamete
- Juvenile: not fully grown or developed
- Adult: fully grown or developed, mature
- Larva: the newly hatched, earliest stage of any of various animals that undergo metamorphosis, differing markedly in form and appearance from the adult
- Postlarva: 1. Stage in the development of a bony fish between the resorption of the yolk sac and the appearance of juvenile characteristics. 2. The larval stage following the nauplius stage in Crustacea, many crustacean groups have a distinct name for this stage
- Spat: an oyster or similar bivalve mollusk in the larval stage, especially when it settles to the bottom and begins to develop a shell
- Subadult: an individual that has passed through the juvenile period but not yet attained typical adult characteristics
- Zoea: a larval form of crabs and other decapod crustaceans, characterized by one or more spines on the carapace and rudimentary limbs on the abdomen and thorax
- Nauplius: the free-swimming first stage of the larva of certain crustaceans, having an unsegmented body with three pairs of appendages and a single median eye
- Polyp: a coelenterate, such as a hydra or coral, having a cylindrical body and an oral opening usually surrounded by tentacles
- Medusa: one of the two forms in which a coelenterate exists. It has a jelly-like umbrella-shaped body, is free swimming and produces gametes
- Ephyra: a stage in the development of discophorous medusae, when they first begin to swim about after being detached from the strobila
- Megalopa: post larva of crustaceans, follows on the zoea
- Planula: free swimming, flattened, ciliated, bilaterally symmetric larval form of various cnidarians, forms from the fertilized egg of a medusa or from a polyp
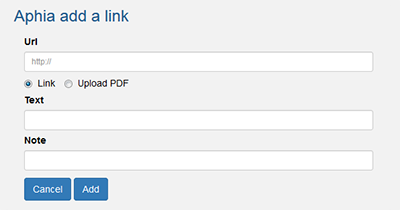
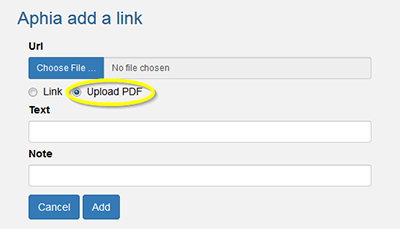
Notes are arranged in three types:
- Checked/added by taxonomic editor or information retrieved through a RAS global species database
- Added by a thematic editor (thematic species databases) (=trusted information)
- Coming from another source (=unreviewed information)
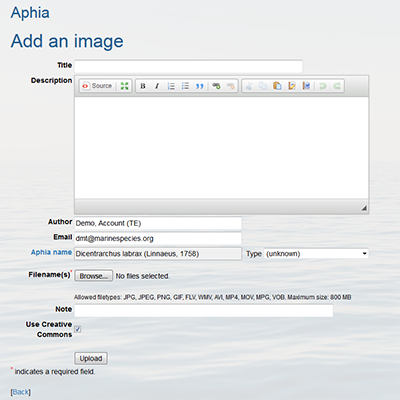
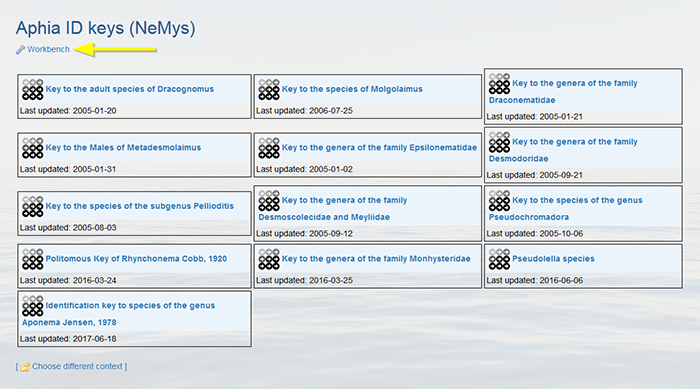
Click on “Workbench”.
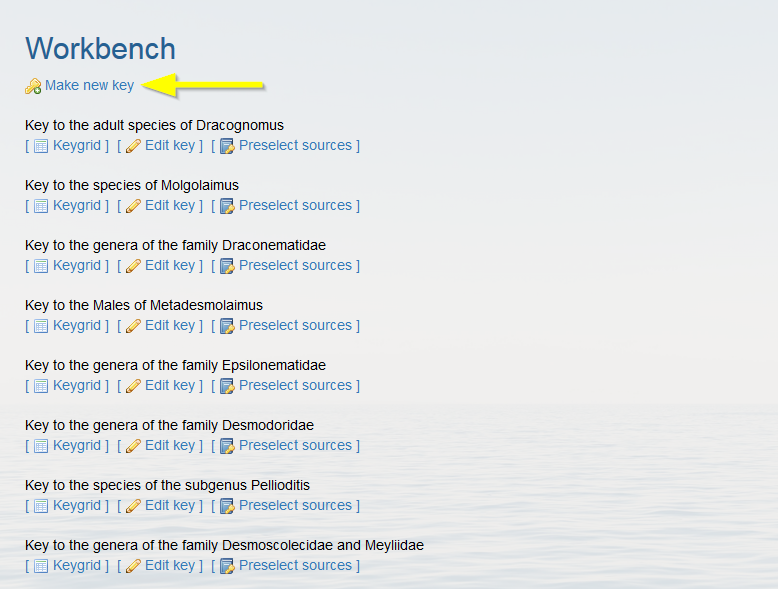
Click on “Make new key”.
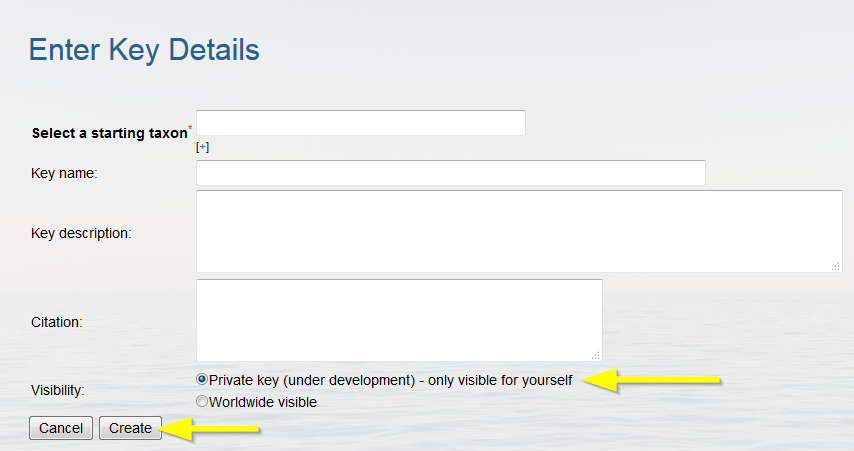
Enter the key details:
- Select a higher starting taxon (e.g. Monhysteridae). Click on [+] if you want to add more taxa.
- Enter the key name. This name will appear in the IDkey overview (e.g. Key to the genera of the family Monhysteridae).
- Enter the description. It is important to mention the reference, or several when it involves an update of an existing key, so that the age of the key and state of update are clear to the users (e.g. Key to the genera of the family Monhysteridae based on the publication of Fonseca and Decraemer (2008)).
- Pick the appropriate visibility. A private key is only visible for yourself and can be published afterwards. A worldwide key is immediately published online.
Click on "Create" when finished.
Your key is now added to the Workbench. Click on [Edit key] to edit the key details and to add the citation. Clicking [Preselect sources] allows you to predefine the sources you will use in your ID Key and easily link these sources to a specific character states (See further). You can efficiently search for datasources by using the search format 'Author%YearOfPublication'. Click on "Find" and on the correct datasource to link the source to your Key.
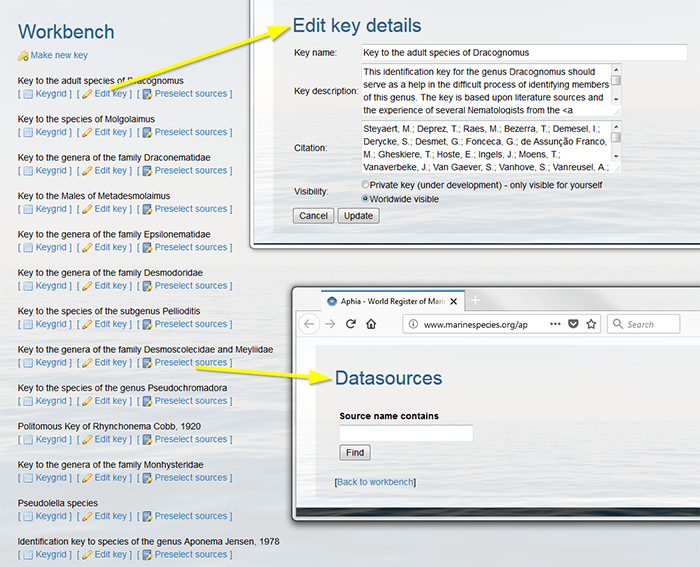
Click on [Keygrid] and “select taxa” to further develop the key.
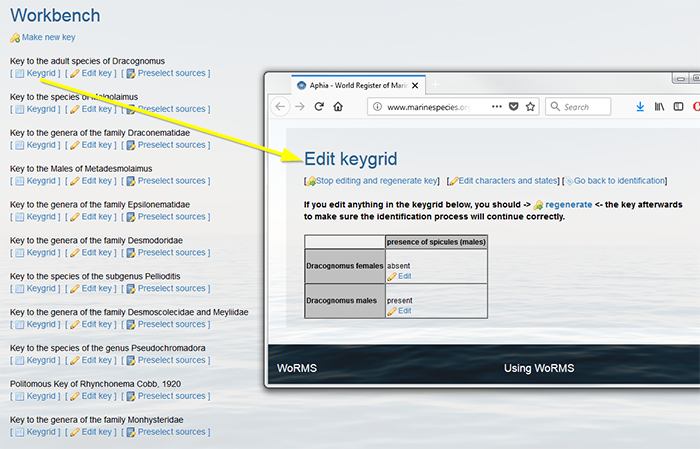
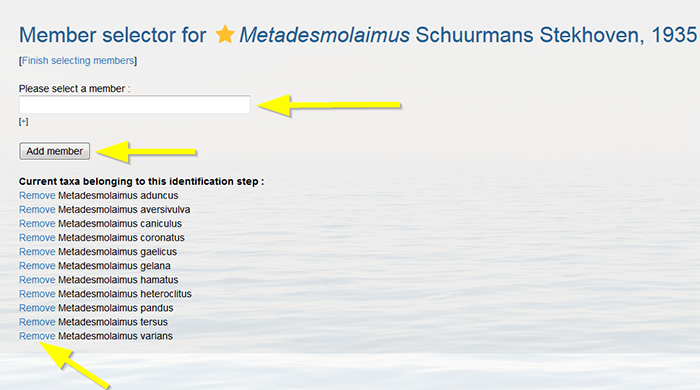
Select every member taxon that you want to include in your key individually. Click on “add member” to upload this new taxon. Note that only taxa available in Aphia can be added. Click on [+] to add more taxa and repeat the latter steps. Click on [Finish selecting members] when all taxa are selected. It is always possible to add more members later.
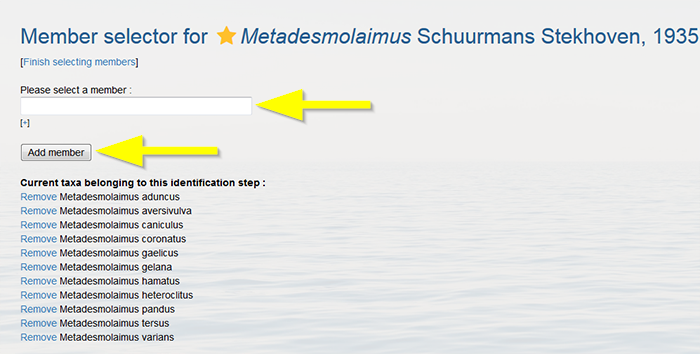
Click on “Edit characteristics and states”. Do not remove the characteristics already displayed.
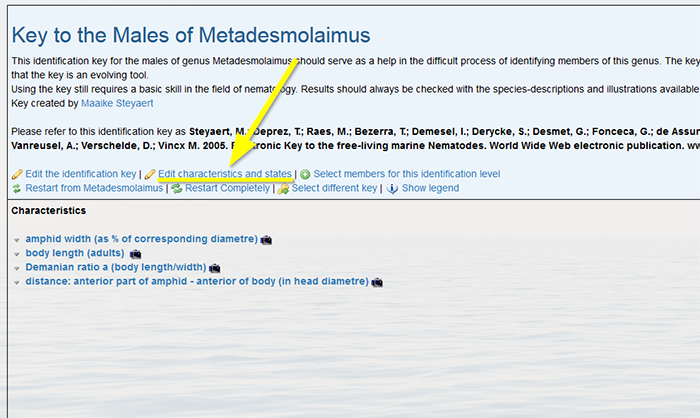
Click again on “[Edit characteristics and states]”. Do not remove the characteristics already displayed.
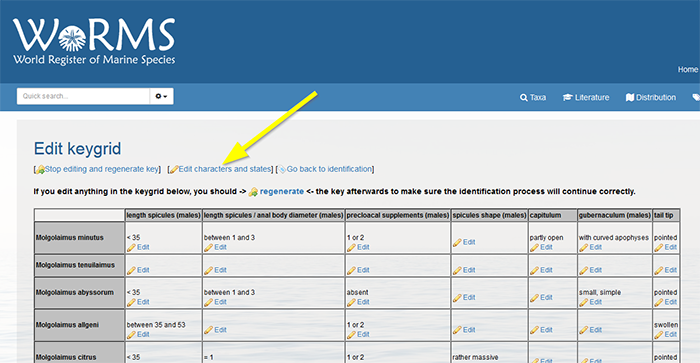
Define the identification characteristics individually in the text box at the bottom of the page. Click on ‘Add characteristic’ to add a new character. Click on the grey arrow left of the character name to define the different states. Fill in the state in the text box and click on ‘Add state’. Add every state separately. The yellow pencil next to the states and characters allows you to upload a drawing or picture.
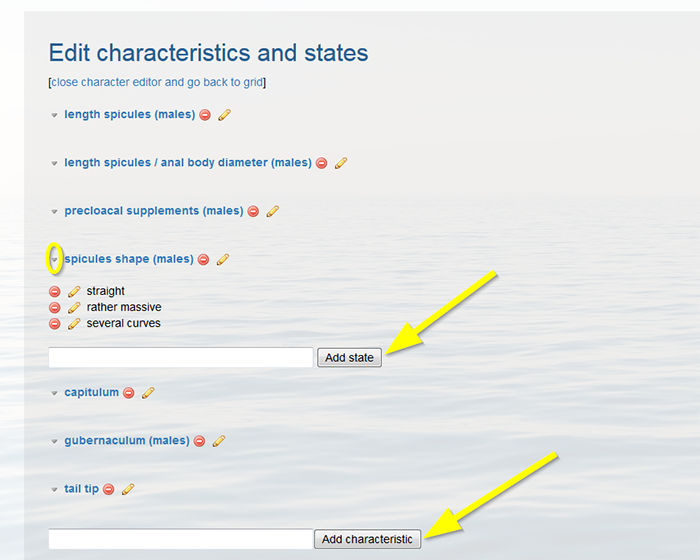
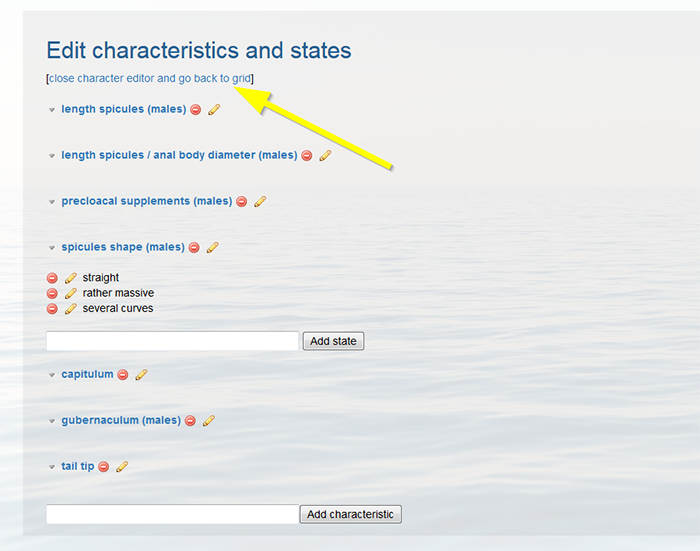
When finished click [close character editor and go back to grid] on top of the page.
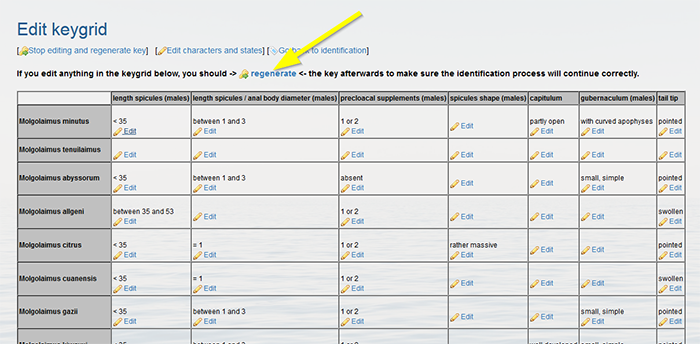
Back in the grid, you can allocate the correct character states to each taxa by clicking “Edit” in the grid cells.
Click on [add state] to assign the correct character state. A character state can be easily removed by clicking [remove character state]. Click on [choose datasource] to link a source.
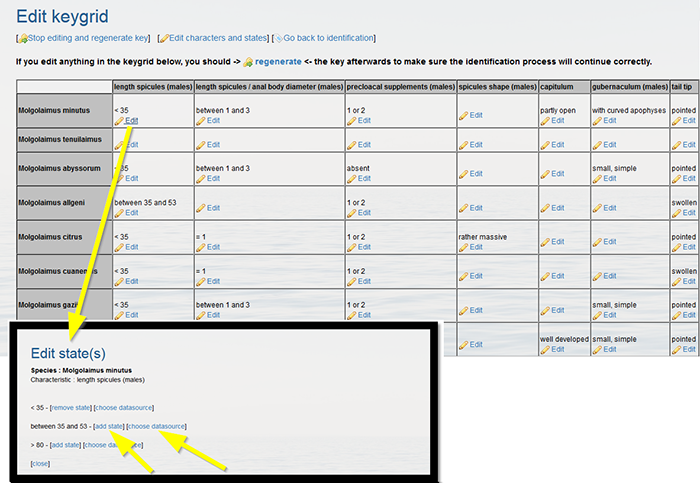
Click "regenerate" when finished.
See also: How can I update the classification?
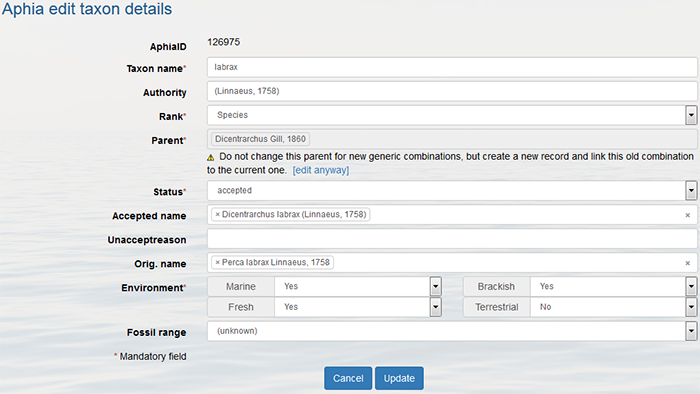
Go to the taxon page and click on [edit taxon]. Change the status into "unaccepted" or any other possible status from the pick list and in case it is a synonym, enter the "accepted name". If possible fill in all other information, for example the unaccept reason and click on [update]. If you change the status of a name, it doesn't mean that the status of its children is changed.
Possible statuses for a name:
- Accepted: Valid name (ICZN) or name considered to be taxonomically correct (ICBN)
- Unreplaced junior homonym: To indicate the unreplaced invalid status of the junior, or later established name, or in the case of simultaneous establishment, the name not given precedence by a first reviser or by an ICZN or ICN ruling
- Nomen protectum: Protected name. A Latin term (meaning "protected name") applied to a name which has been given precedence over its unused senior synonym or senior homonym relegated to the status of nomen oblitum (q.v., and see Article 23.9.2).
- Nomen novum: New replacement name. A new name published as an explicit substitute (avowed substitute) for a legitimate or illegitimate, previously published name, which is its replaced synonym and which, when legitimate, does not provide the final epithet, name, or stem of the replacement name (Art. 6.11 and 7.4; for names not explicitly proposed as substitutes see Art. 6.12 and 6.13).
- Unaccepted: Synonym name, or anything that is not accepted
- Nomen nudum: A Latin term referring to a name that, if published before 1931, fails to conform to Article 12; or, if published after 1930, fails to conform to Article 13. A nomen nudum is not an available name, and therefore the same name may be made available later for the same or a different concept; in such a case it would take authorship and date [Arts. 50, 21] from that act of establishment, not from any earlier publication as a nomen nudum.
- Interim unpublished: An as yet unavailable name (until in a print issue) which has been published online only, in a work that does not show evidence of ZooBank registration (ICZN Article 8.5)
- Superseded combination: When a species is transferred to another genus, and is also known as a new combination
- Junior homonym: To indicate the invalid status of the junior, or later established name, or in the case of simultaneous establishment, the name not given precedence by a first reviser or by an ICZN or ICN ruling. This is known as a later homonym in the ICN.
- Junior subjective synonym: Applied as the reason for the unaccepted status for the junior, or later established, subjective synonym (a published opinion that two names apply for the same taxon). This is known as an heterotypic synonym in the ICN.
- Junior objective synonym: Applied to the later established name in cases where two or more scientific names have the same type material. This is known as a homotypic synonym in the ICN.
- Nomen oblitum: A Latin term (meaning ‘forgotten name’) applied after 1 January 2000 to a name, unused since 1899, which as a result of an action taken under ICZN Article 23.9.2 does not take precedence over a younger synonym or homonym in prevailing usage
- Misspelling - incorrect original spelling: An error in the name's original publication that doesn't comply with ICZN naming rules. Corrections to these errors are termed "justified emendations" by the ICZN, requiring compliance with ICZN rules and guidelines (see section 8 in Welter-Schultes 2012)
- Misspelling - incorrect subsequent spelling: Erroneous name in the literature that need to be captured, e.g., when widespread, or needed in the database
- Unjustified emendation: Nomenclatural act under ICZN article 33.2 and must strictly be a ‘demonstrably intentional change’ of the original spelling
- Incorrect grammatical agreement of specific epithet: Mandatory change in spelling consequent upon changes in rank or combination. See Welter-Schultes (2012) section 8.1.6
- Misapplication: When a name has been misapplied to an incorrectly identified specimen.
- Unavailable name: Not compliant with the relevant ICZN or ICN code. See also How to deal with unavailable names?
- Superseded rank: When a subspecies, subgenus, subfamily, etc. is moved up or down within the hierarchy (i.e., subspecies now accepted as a species or vice versa)
- Nomen rejiciendum: (1) A name which, under the provisions of the Code, cannot be used as a valid name and which is set aside in favour of another name. (2) A name which, as a matter of taxonomic judgment, is either treated as a junior subjective synonym (q.v.) of a name used as valid or is believed not to be applicable to the taxon under consideration.
- Alternative representation: An accepted name with (or without) a subgenus, but slightly less preferred. See also How to deal with species with accepted subgenera using the status of alternative representation?
- Temporary name: Ad-hoc higher rank taxa of convenience to accommodate child taxa for which the classification is not yet finalised. i.e. incertae sedis, sp. a, ...
- Uncertain: To indicate taxonomic or nomenclatural uncertainty for cases which cannot be classed as either 'accepted' or 'unaccepted'
- Nomen dubium: A name of uncertain taxonomic significance (no type and/or original description very vague)
- Taxon inquirendum: A taxon, of which the taxonomic validity is uncertain or disputed by different experts (unable to tell what it is)
- Unassessed: Name coming from nomenclator (lists of names) or from, e.g., museum collection database, where the status might be known in the published literature, but for which the status has not yet been researched, assessed and documented
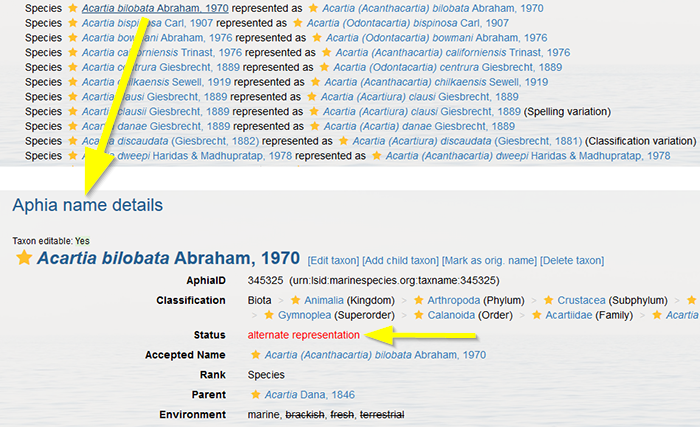
Each record status carries a quality indicator:
When you have changed taxonomic information of a taxon, the record status will automatically be set to "checked: verified by taxonomic editor", represented by the fully filled yellow start. The edit history will include "date" and "changed" by "your name".
When no changes are required you can validate the taxon by clicking on the quality indicator pictogram, the record status will be set to "checked: verified by taxonomic editor".
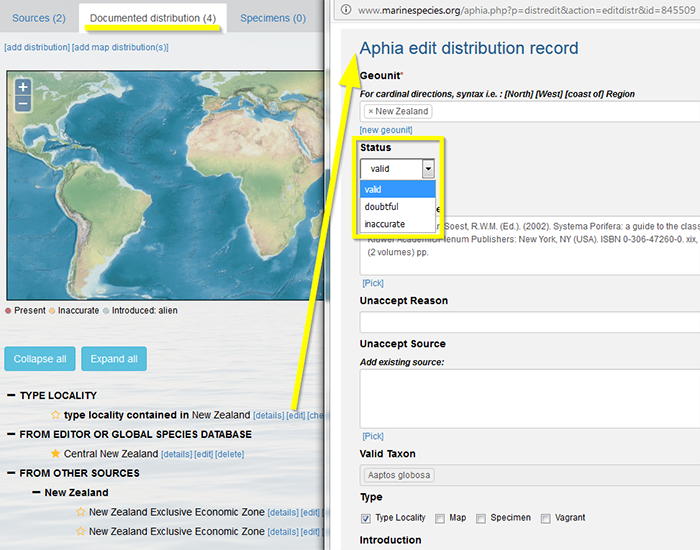
Top Change (the status of) a distributionGo to the taxon details page and click on [edit] next to the distribution record. There is a drop down list with 3 possibilities:
- Valid: the record is certainly valid
- Doubtful: doubts on whether the species really lives there
- Inaccurate: taxon was cited for this area but it is clear that it does not live there (misidentification, wrong label, etc)
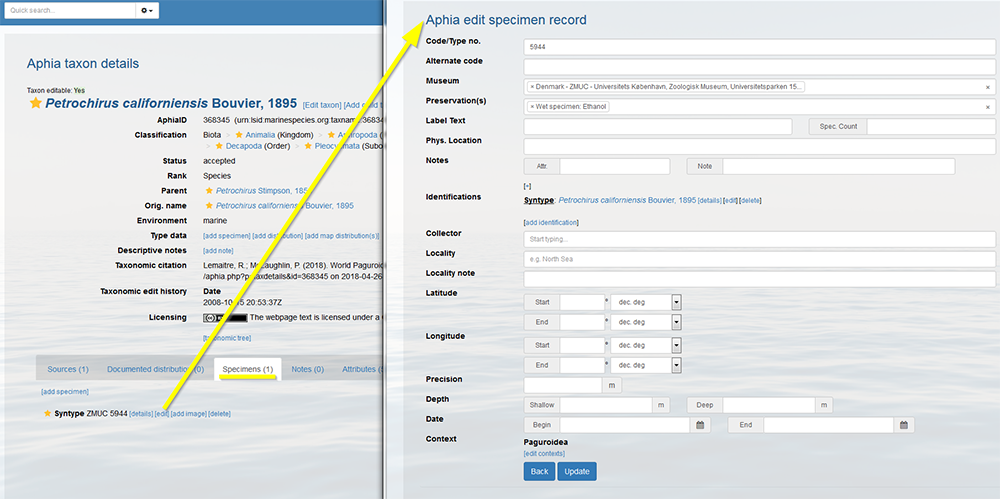
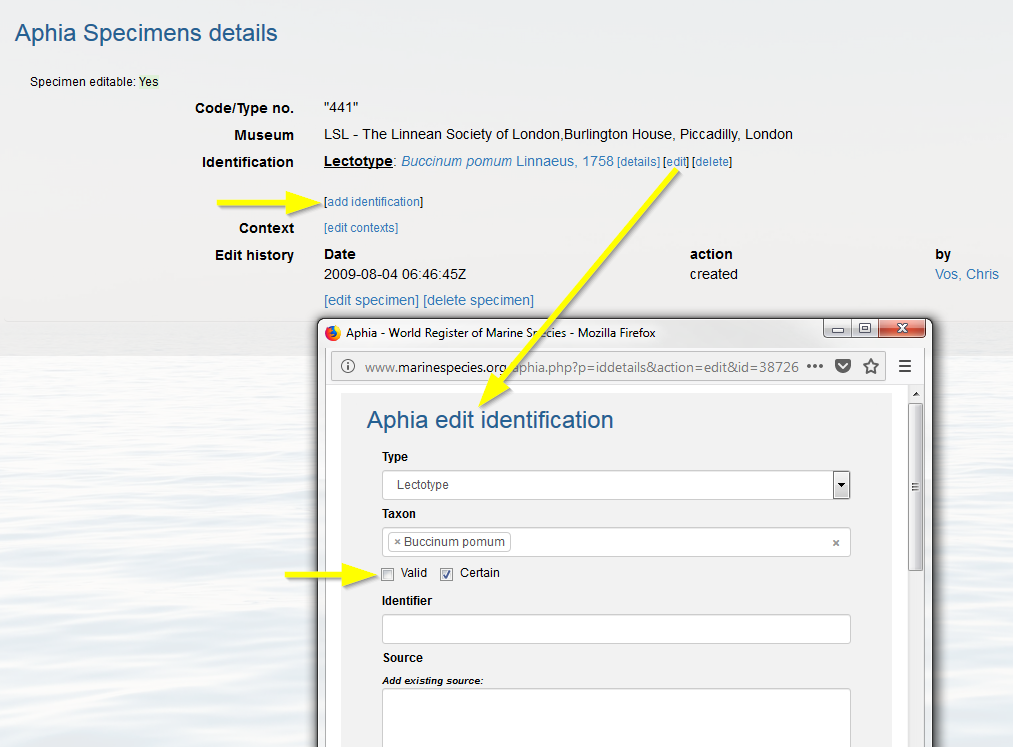
If a specimen description has been published, but afterwards turns out to be given the wrong name, you can create a trace of its identification history. Go to the taxon page and click on [details] or [edit] next to the specimen record and click on [add identification]. Do not forget to edit the previous identification by flagging-off the valid button.
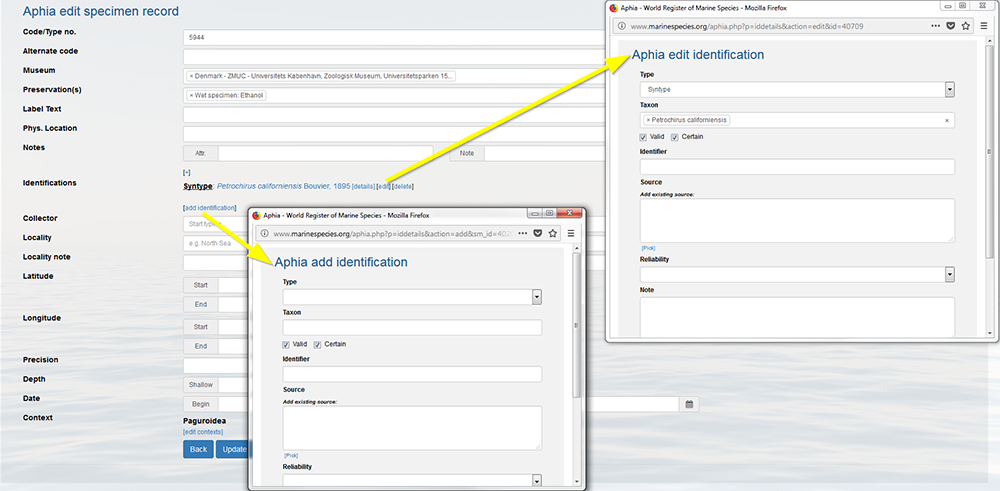
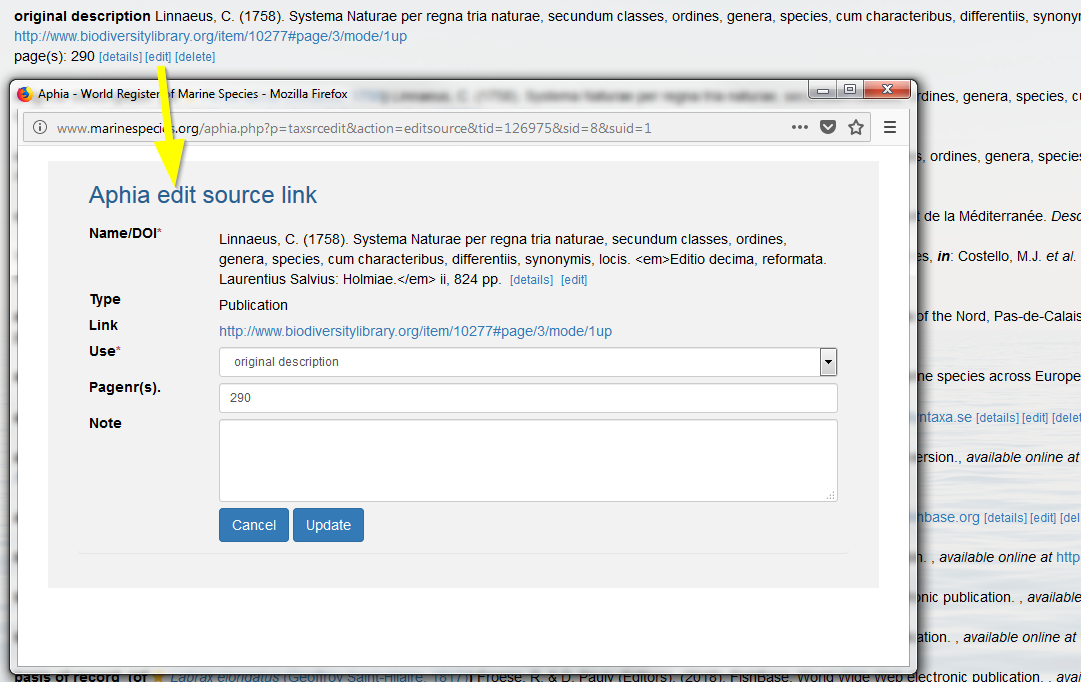
If however you want to correct the source, e.g. the spelling of the reference, you have to go to the source details in the literature module, and subsequently click on [edit source]. The changes are automatically reflected throughout the entire database.
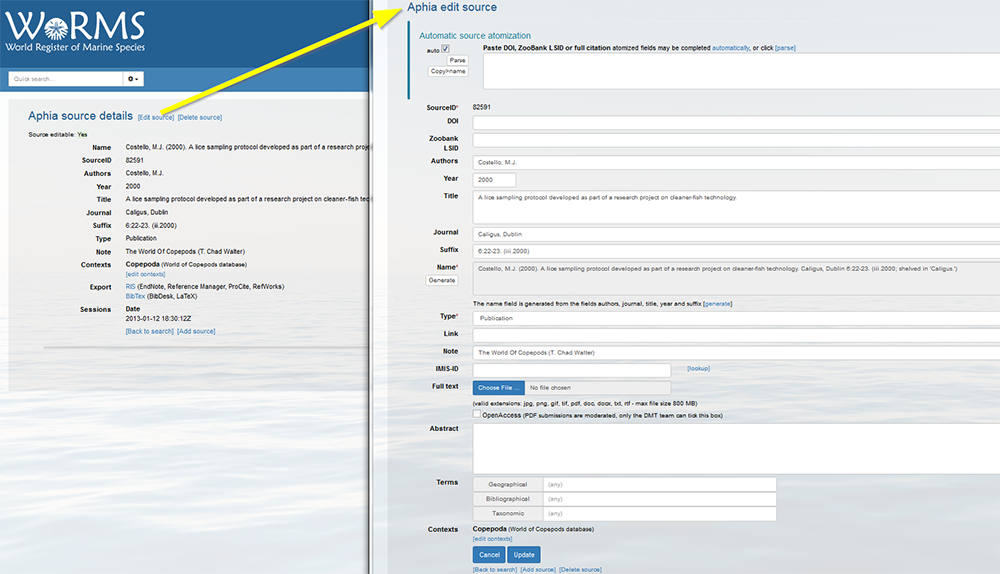
Log in to the database, go to [My Aphia] and click on "Similar sources (de-duplication tool)".
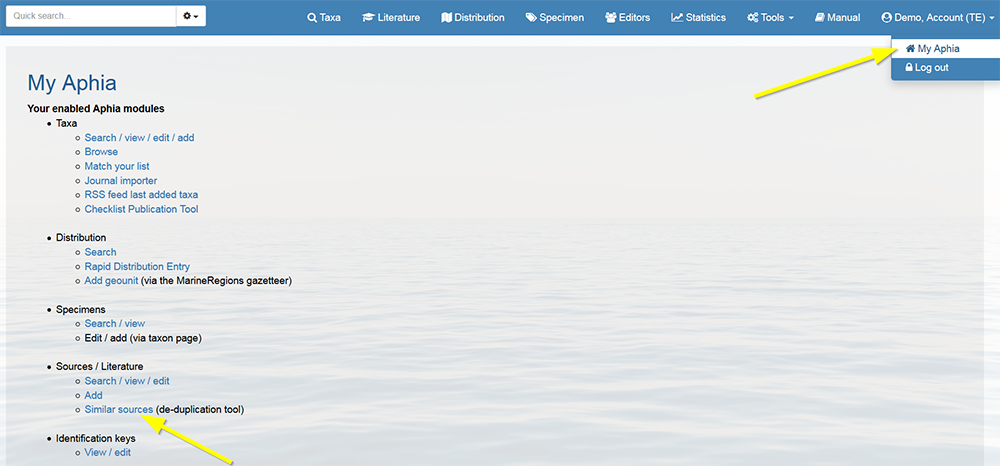
You will see a list of possible duplicate sources. The list is personalized, depending on your edit rights.
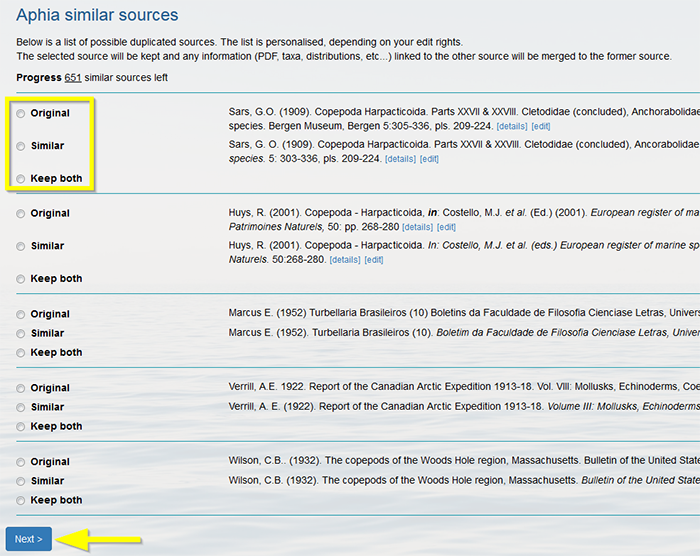
For each source, you can choose to keep the original source, the similar source or both (in case of non-duplicates). If you click on "Next" the selected source will be retained in the database and any information (PDF, taxa, distributions, etc.) linked to the other source will be merged to the retained source. If you choose to keep both, both sources will be retained in the database as separate sources and no information will be merged. Top Journal importer This feature allows you to semi-automatically import (newly) published names coming from ZooBank and Pensoft.
Log in to the database, go to [My Aphia] and click on "Journal importer".
By pasting either a Pensoft DOI or a ZooBank Reference LSID (containing Nomenclatural Acts), the source and the taxon/taxa will be imported in a "guided" way.
Supply a Pensoft DOI (e.g. 10.3897/zookeys.449.8544) or ZooBank Reference LSID (e.g. urn:lsid:zoobank.org:pub:CDE9DC2E-0EF0-4787-895C-2B645470D2B5) and click on "Lookup".
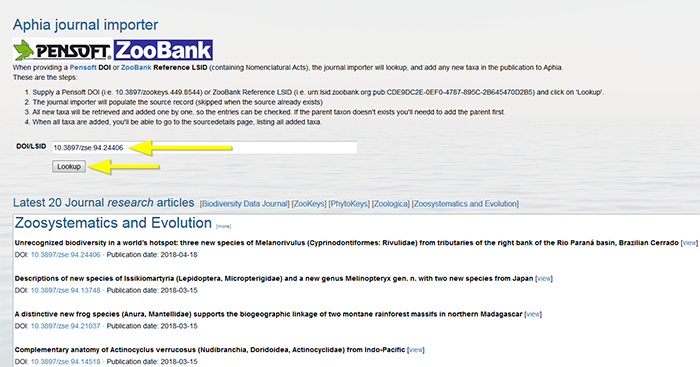
The journal importer will populate the source record (skipped when the source already exists). Verify if all information is correct. Click on "Next".
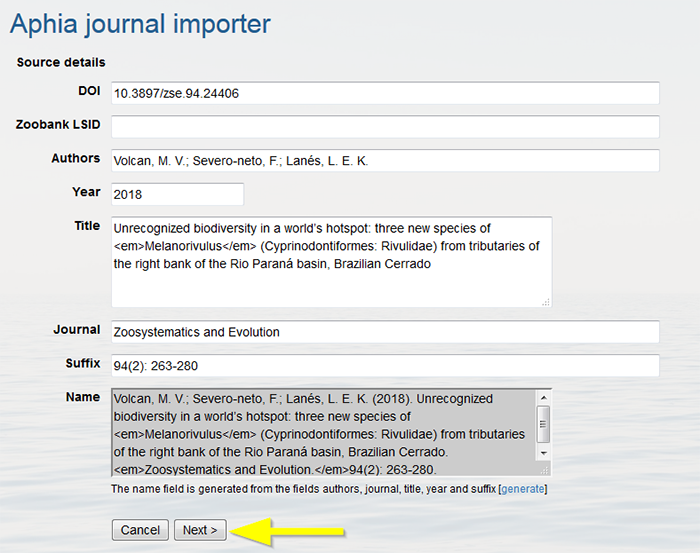
All new taxa will be retrieved and added one by one, so the entries can be checked. If the parent taxon is not available you will need to add the parent first. Mark the environment(s) in which the taxon occurs (environment is now a mandatory field when adding new species), and click "Next".
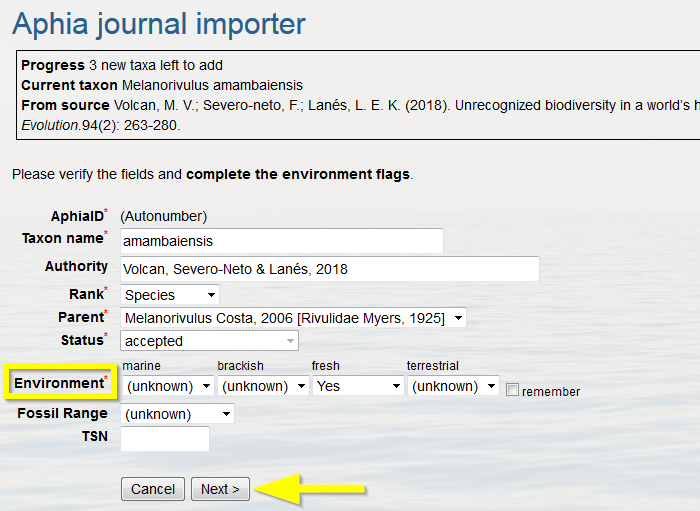
When all taxa are added, you will be able to go to the source details page, listing all added taxa.
The latest Pensoft journals publications are also listed, and through one click on the DOI, one can start importing the chosen publication.
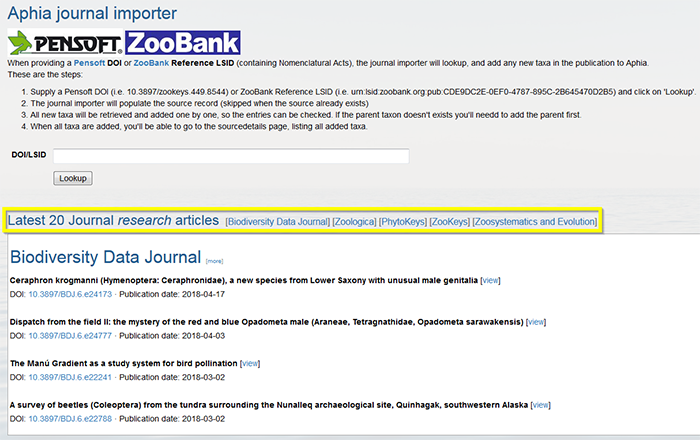
A demonstration video of the Aphia journal importer can be found here. Top Checklist Publication Tool This feature allows you to create a taxonomic checklists on-the-fly. It may help editors to prepare a checklist for publication.
Log in to the database, go to [My Aphia] and click on "Checklist Publication Tool".
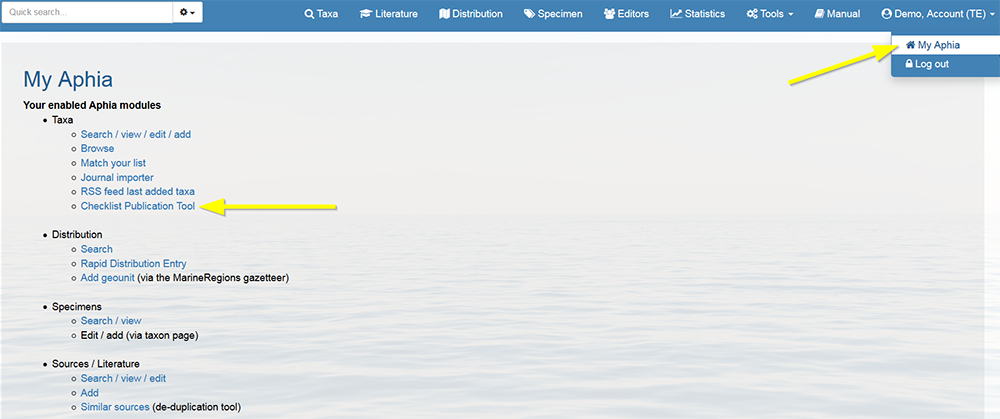
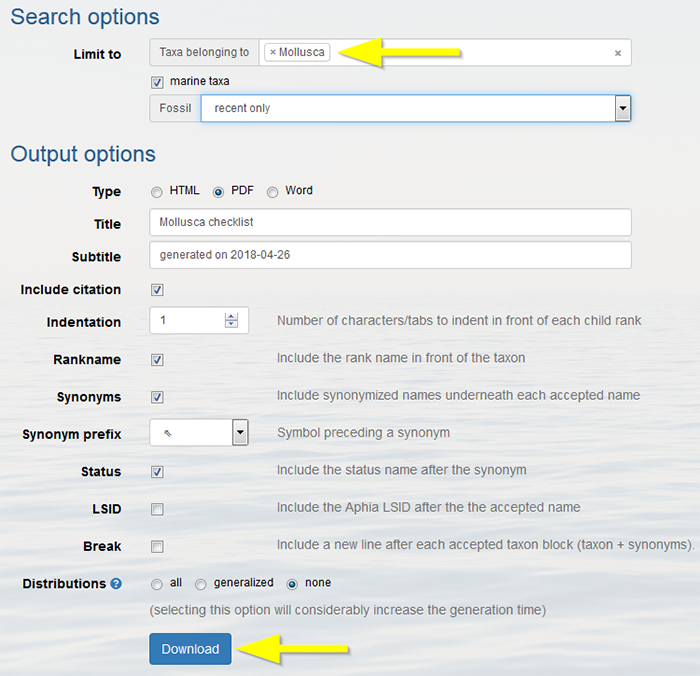
Select the taxon for which you want to create a checklist (e.g. Mollusca). After choosing all the output options, click on "Download" . The checklist can be downloaded as a html, PDF or Word document.
In principle we do not delete information which has been published in primary literature or which is in common use (e.g. many hits in google). In that case you need to correct the error by changing the status (see Edit taxon details).
Sometimes, information can be wrongly entered into the database or spelling mistakes were made. Sometimes entries into the database are based on published spelling mistakes, but the name never got in common use. This kind of information should be removed as soon as possible before they start their own life by either correcting the alternate representation or deletion.
To remove information from the database, you need to click on the respective [delete] buttons. An email is sent to the database management team, who can finally delete the record. When this is done they will inform you via email.
Note that each record has a unique AphiaID. The AphiaIDs of the deleted items are never overwritten. The reason of the deletion of a record is archived. This is important for other data systems that are linked to these records.
TopFrequently asked questions
How come there is a difference between RAS and some of the contexts like Isopoda etc ?It is important to understand that all the information in RAS is in a single database (so each name has a single instance!), but on the basis of contexts (compare it with 'tags'), we can make collections (contexts) in RAS and display it on different web interfaces. For Editors, the context option is available near the bottom of each edit form. If you add new information to RAS and you don’t want to be bothered flagging on the context to each new item, then login via your own section website (same login) and your context is automatically added to every new record.
Top How can I know what species of my taxon are in RAS, but are not part of my collection? Go to the search tool, and select your taxon in the box "Limit to taxa belonging to" and select your collection context in the "Exclude context" box. You can further choose the status from a drop-down list and you can limit the search to non checked taxa or taxa that are in quarantine by flagging off the right box. You can also specify environment, taxon rank, etc …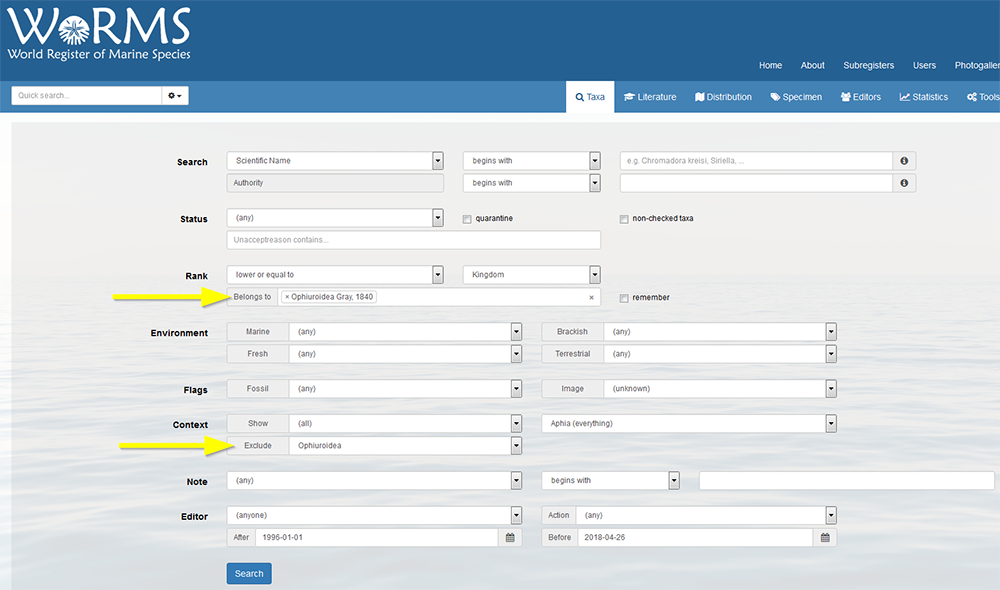
Go to the search tool, and select the action (all, created, changed or checked), indicate the person and if necessary select a begin date. You can further set other items such as limit to species rank, limit to a particular family or valid names only, etc...
For information added in the last week see the Weekly Digest
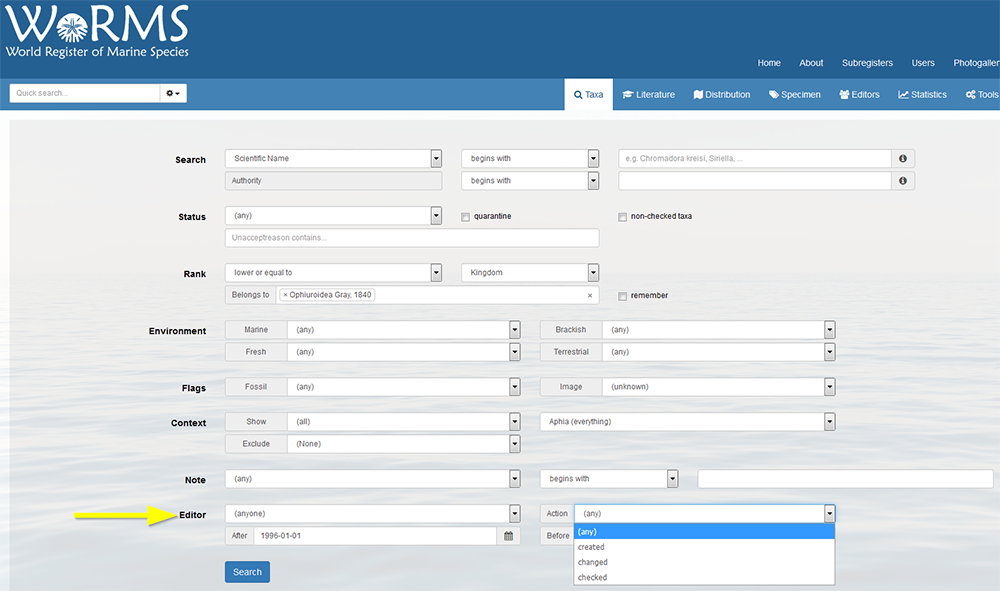
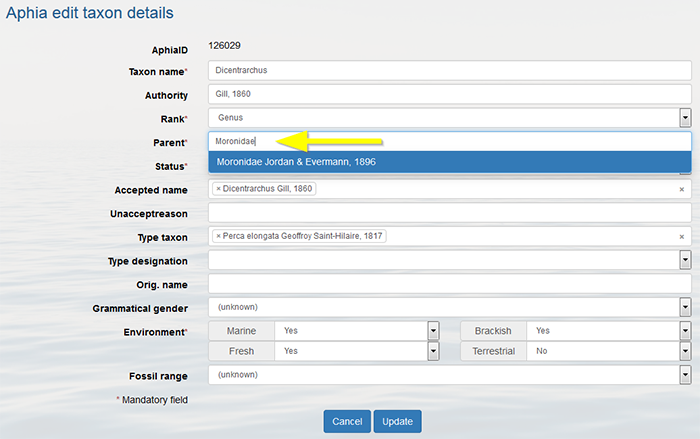
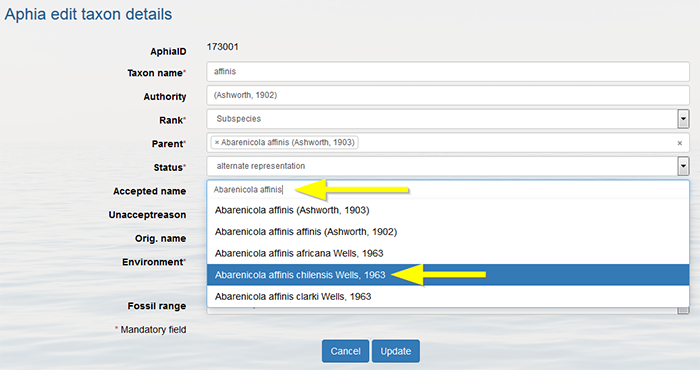
You can only add a subspecies if the parent species is present in the database. Use the status "alternate representation" for the species and link it to its nominal subspecies in case the species name is not used separately.
It often happens that a higher taxon became invalid and its children are spread over many other taxa. When you know what the type taxon is, then the accepted taxon is the new parent of the type taxon. The problem is of course when you don't know what the type taxon is. In that case you have to contact the data management team here and ask to change the status without linking to an accepted taxon. You always have to add the reason of the invalid status.
Top How do I enter an occurrence record in a paper that mentions the original paper, which also used a synonym name?The ideal solution would be to have the possibility to link a single distribution record to more than one name and source. But the system is not built to cope with this so we suggest you enter the information as it was published in the paper that you are encoding, but that you add a note citing the original reference and original name.
Top What should I do if not all genera can be assigned to a subfamily for example?If for example there is a taxon (any rank) that doesn't fit anywhere in the current classification, you must create a temporary parent with the name "X incertae sedis", where X is the name of the first following parent.
Some people would like to skip the sub or super or infra ranks in the classification. We leave this decision up to the editor.
See examples at: https://ras.biodiversity.aq/aphia.php?p=taxlist&tName=%incertae%20sedis .
Top Can we emphasize preferred vernacular names?Yes this is possible. When adding/editing a vernacular name you can tick the "preferred" box. However, trying to settle on a single "correct" common name is tricky. Developing a complex thesaurus of common names falls more within the scope of an ethnographic or linguistic project. In many cases the vernacular names come from the older literature and we should also record these in RAS for the benefit of people working with that old material.
Top Can I add taxa that are in press?In principle we do not add taxa to RAS that have not been formally published according to the different Codes. Only if your paper is in press, so not before submission and only after acceptance, you may add the taxa to RAS or change the status of taxa. But always clearly refer to the source by linking to the reference of your paper. This also counts for synonyms. If you think the species is a synonym, but you have not published it, you can add a note like this: "The validity of this recently described species needs to be re-evaluated".
Top Can you explain how the citation on each page works?In the same way as citing regular publications we would like to encourage citing data that have been published online. When a taxon name has been created, changed or marked as checked by an editor, he or she will automatically become the (co)author of that webpage. The citation has the format as a chapter in a book: [author(s)] [Year]. [chapter] in [Book]. [Publisher]. [Page]
=> Editors (Year). Taxon + Authority. In: GSD title. Accessed through the World Register of Marine Species at URL on [date].
When the taxon has not yet been verified by an expert, RAS is listed as the author. A member of the data management team is not considered an expert of the taxon and therefore will not receive "the authority" for a webpage.
Top When adding a specimen, should I use the original species name or the currently accepted?It is better to add the specimen details to the original species name. So you don't loose information, and it is easier to trace back the specimen in the museum collection. Besides, because the taxon name is linked to the currently accepted name, the information on the specimen is also shown on the webpage of accepted taxon.
Top How are we dealing with the correct citation of the year of publication of a new species´ description when the online version and the printed version appear in different years?When you cite the publication containing the original description, you cite it with the date of the print edition, but also include the date of the online edition in the original description reference, so people can see that there is no oversight from the part of the RAS editor.
For example: Neopycnodonte zibrowii
Gofas, Salas & Taviani in Wisshak, Lopez Correa, Gofas, Salas, Taviani, Jakobsen & Freiwald, 2009
The source says: Deep Sea Research I 56(3): 374-404 [published online 29 October 2008; printed copies
deposited by January 2, 2009, in several museum libraries as stated in acknowledgements; printed edition of
the journal dated March 2009].
Art. 5.3 "A typographical sign such as ? [...], when used to qualify the application of a scientific name, does not form part of the name of a taxon [...]. So, you just drop the question mark.
Top How should I handle species that currently are *tentatively* placed in synonymy (the kind of thing that is usually preceded by a "?" in synonymies)?Flag them as synonyms, and under "unaccepted reason" write "uncertain synonym".
Top How can I export a source in RIS and BibTex format? All sources that have been atomized can be exported to the RIS (EndNote, Reference Manager, ProCite, RefWorks) and BibTex (BibDesk, LaTeX) formats, including the search result. These export links can be found on the bottom of each source record.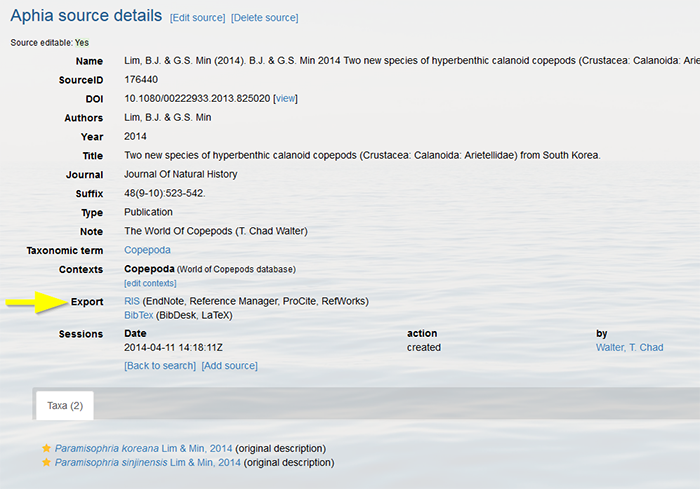
Literature published since 1923 is not in all cases Open Access. When editors upload pdfs for post-1923 publications, they will initially show as Non Open Access.
Regularly, the DMT screens these publications and - in collaboration with the VLIZ library - it is verified which of these publications are Open Access, and can thus be offered to the users as such. As this is a manual recurring task, it can still happen that we might have missed some publications in putting them Open Access. If that would be the case, don't hesitate to contact us, and provide us the SourceID(s) of these publications/journals. Top Where should I place unaccepted genera in the taxonomic hierarchy? It is recommended that genera which are no longer accepted and are synonymised with another genus should be placed in the taxonomic hierarchy in the family to which the type-species of the synonymised genus belongs - no matter in which family these genera were originally placed. Top How to deal with unavailable names? Some journals currently pose a problem in terms of the validity of published names. In cases of online only journals (where there is no printed or only limited distribution of the journal), and where no Zoobank registration of the name is apparent, then the taxon is not validly published according to the ICZN (ICZN Art. 8.5.3), and is an unavailable name.
Names from such journals should be added to WoRMS with the status ‘unaccepted > unavailable name’. The original source of the name should be added as ‘original description (unavailable nomenclaturally)’. There should be no ‘original description’ or ‘original name’ entry for these names.
In cases like this, the authors of the taxon should be contacted and should rectify this situation, so the name becomes validly published. When the new publication which validates the name is published, then that validly published name should be entered into WoRMS with the new publication linked as the ‘original description’. The unavailable name can be then be linked to the newly accepted name and will appear as a ‘synonymised name’. The valid publication source can also be linked to the unavailable name as a ‘status source’. The names may have different publication years, and possibly even different authorships (as for the second example given here).
Here are two examples which indicate how such names should be entered into WoRMS:
- Stenothoe bella Krapp-Schickel & Lo Brutto, 2015
- Stenothoe bella Krapp-Schickel & Lo Brutto, 2015, was not validly published (online only, not registered with Zoobank), so is entered as an unavailable.
- The authors were contacted and validly published the name as Stenothoe bella Krapp-Schickel & Lo Brutto, 2022.
- The names are linked, sources are provided, and there are explanatory notes attached where relevant.
- Eotiaris guadalupensis Thompson in Thompson, Petsios, Davidson, Erkenbrack, Gao & Bottjer, 2015 †
- Eotiaris guadalupensis Thompson in Thompson, Petsios, Davidson, Erkenbrack, Gao & Bottjer, 2015 † was published online only and not registered in Zoobank.
- It was published validly two years later: Eotiaris guadalupensis Thompson in Thompson, Petsios & Bottjer, 2017 †
Both "Genus species" and "Genus (Subgenus) species" are valid (accepted) names for the same species (e.g. Acartia japonica Mori, 1940).
From the DMT viewpoint, "Genus (Subgenus) species" is the accepted name. The other name, "Genus species", is also an accepted name but is less preferred and should be indicated as an "alternative representation". Only one can be marked as "accepted" to avoid double-counting of species in the statistics. This preference may vary depending on taxonomic context or editorial decision (e.g. Alvania (Linemera) beyersi (Thiele, 1925)).
Case 1: Only "Genus (Subgenus) species" is listed in WoRMS
- Add "Genus species" as a separate name.
- Link "Genus species" to "Genus (Subgenus) species" as an 'alternative representation'.
Case 2: Only "Genus species" and "Genus (Subgenus)" are in WoRMS
- Add "Genus (Subgenus) species" as a new entry.
- Link "Genus species" to "Genus (Subgenus) species" as an 'alternative representation'.
When the subgenus is not valid (for any reason): "Genus species" and "Genus (Subgenus) species" are not equivalent. The combination using the invalid subgenus ("Genus (Subgenus) species") should be marked as 'Unaccepted' – not an alternative representation (e.g. Trochus (Lischkeia) P. Fischer, 1879).